SARS-CoV-2 N protein promotes NLRP3 inflammasome activation to induce hyperinflammation
- PMID: 34341353
- PMCID: PMC8329225
- DOI: 10.1038/s41467-021-25015-6
SARS-CoV-2 N protein promotes NLRP3 inflammasome activation to induce hyperinflammation
Erratum in
-
Author Correction: SARS-CoV-2 N protein promotes NLRP3 inflammasome activation to induce hyperinflammation.Nat Commun. 2021 Aug 31;12(1):5306. doi: 10.1038/s41467-021-25629-w. Nat Commun. 2021. PMID: 34465791 Free PMC article. No abstract available.
Abstract
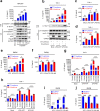
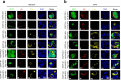
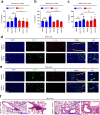
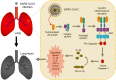
Excessive inflammatory responses induced upon SARS-CoV-2 infection are associated with severe symptoms of COVID-19. Inflammasomes activated in response to SARS-CoV-2 infection are also associated with COVID-19 severity. Here, we show a distinct mechanism by which SARS-CoV-2 N protein promotes NLRP3 inflammasome activation to induce hyperinflammation. N protein facilitates maturation of proinflammatory cytokines and induces proinflammatory responses in cultured cells and mice. Mechanistically, N protein interacts directly with NLRP3 protein, promotes the binding of NLRP3 with ASC, and facilitates NLRP3 inflammasome assembly. More importantly, N protein aggravates lung injury, accelerates death in sepsis and acute inflammation mouse models, and promotes IL-1β and IL-6 activation in mice. Notably, N-induced lung injury and cytokine production are blocked by MCC950 (a specific inhibitor of NLRP3) and Ac-YVAD-cmk (an inhibitor of caspase-1). Therefore, this study reveals a distinct mechanism by which SARS-CoV-2 N protein promotes NLRP3 inflammasome activation and induces excessive inflammatory responses.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

