NR4A family members regulate T cell tolerance to preserve immune homeostasis and suppress autoimmunity
- PMID: 34343134
- PMCID: PMC8492309
- DOI: 10.1172/jci.insight.151005
NR4A family members regulate T cell tolerance to preserve immune homeostasis and suppress autoimmunity
Abstract
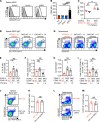
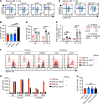
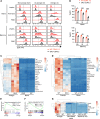
The NR4A family of orphan nuclear receptors (Nr4a1-3) plays redundant roles to establish and maintain Treg identity; deletion of multiple family members in the thymus results in Treg deficiency and a severe inflammatory disease. Consequently, it has been challenging to unmask redundant functions of the NR4A family in other immune cells. Here we use a competitive bone marrow chimera strategy, coupled with conditional genetic tools, to rescue Treg homeostasis and unmask such functions. Unexpectedly, chimeras harboring Nr4a1-/- Nr4a3-/- (double-knockout, DKO) bone marrow developed autoantibodies and a systemic inflammatory disease despite a replete Treg compartment of largely WT origin. This disease differs qualitatively from that seen with Treg deficiency and is B cell extrinsic. Negative selection of DKO thymocytes is profoundly impaired in a cell-intrinsic manner. Consistent with escape of self-reactive T cells into the periphery, DKO T cells with functional, phenotypic, and transcriptional features of anergy accumulated in chimeric mice. Nevertheless, we observed upregulation of genes encoding inflammatory mediators in anergic DKO T cells, and DKO T cells exhibited enhanced capacity for IL-2 production. These studies reveal cell-intrinsic roles for the NR4A family in both central and peripheral T cell tolerance and demonstrate that each is essential to preserve immune homeostasis.
Keywords: Autoimmune diseases; Autoimmunity; Immunology; T cell development; Tolerance.
Conflict of interest statement
Figures