Evolution of glutamatergic signaling and synapses
- PMID: 34343611
- PMCID: PMC9233959
- DOI: 10.1016/j.neuropharm.2021.108740
Evolution of glutamatergic signaling and synapses
Abstract
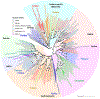
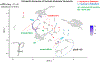
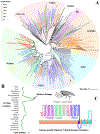
Glutamate (Glu) is the primary excitatory transmitter in the mammalian brain. But, we know little about the evolutionary history of this adaptation, including the selection of l-glutamate as a signaling molecule in the first place. Here, we used comparative metabolomics and genomic data to reconstruct the genealogy of glutamatergic signaling. The origin of Glu-mediated communications might be traced to primordial nitrogen and carbon metabolic pathways. The versatile chemistry of L-Glu placed this molecule at the crossroad of cellular biochemistry as one of the most abundant metabolites. From there, innovations multiplied. Many stress factors or injuries could increase extracellular glutamate concentration, which led to the development of modular molecular systems for its rapid sensing in bacteria and archaea. More than 20 evolutionarily distinct families of ionotropic glutamate receptors (iGluRs) have been identified in eukaryotes. The domain compositions of iGluRs correlate with the origins of multicellularity in eukaryotes. Although L-Glu was recruited as a neuro-muscular transmitter in the early-branching metazoans, it was predominantly a non-neuronal messenger, with a possibility that glutamatergic synapses evolved more than once. Furthermore, the molecular secretory complexity of glutamatergic synapses in invertebrates (e.g., Aplysia) can exceed their vertebrate counterparts. Comparative genomics also revealed 15+ subfamilies of iGluRs across Metazoa. However, most of this ancestral diversity had been lost in the vertebrate lineage, preserving AMPA, Kainate, Delta, and NMDA receptors. The widespread expansion of glutamate synapses in the cortical areas might be associated with the enhanced metabolic demands of the complex brain and compartmentalization of Glu signaling within modular neuronal ensembles.
Keywords: Algae; Aplysia; Aspartate; Cnidaria; Ctenophores; Eukaryotes; GABA; Glutamate receptors; Glutamine; Nervous system evolution; Neurotransmitters; Placozoa; Stress; Synapse; Trichoplax; Vesicular glutamate transporters; scRNA-seq.
Copyright © 2021 Elsevier Ltd. All rights reserved.
Conflict of interest statement
Declaration of competing interest
The authors have no conflict of interest to declare.
Figures










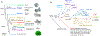



References
-
- Abelson PH, 1959. Geochemistry of organic substances. In: Abelson PH (Ed.), Researches in Geochemistry. Wiley, New York, pp. 79–103.
-
- Alfieri A, Doccula FG, Pederzoli R, Grenzi M, Bonza MC, Luoni L, Candeo A, Romano Armada N, Barbiroli A, Valentini G, Schneider TR, Bassi A, Bolognesi M, Nardini M, Costa A, 2020. The structural bases for agonist diversity in an Arabidopsis thaliana glutamate receptor-like channel. Proc. Natl. Acad. Sci. U. S. A 117, 752–760. - PMC - PubMed
-
- Anctil M, Carette JP, 1994. Glutamate immunoreactivity in non-neuronal cells of the sea anemone Metridium senile. Biol. Bull 187, 48–54. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

