Concatemeric Broccoli reduces mRNA stability and induces aggregates
- PMID: 34347781
- PMCID: PMC8336797
- DOI: 10.1371/journal.pone.0244166
Concatemeric Broccoli reduces mRNA stability and induces aggregates
Abstract
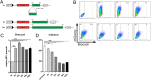
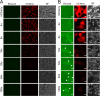
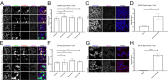
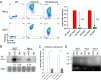
Fluorogenic aptamers are an alternative to established methodology for real-time imaging of RNA transport and dynamics. We developed Broccoli-aptamer concatemers ranging from 4 to 128 substrate-binding site repeats and characterized their behavior fused to an mCherry-coding mRNA in transient transfection, stable expression, and in recombinant cytomegalovirus infection. Concatemerization of substrate-binding sites increased Broccoli fluorescence up to a concatemer length of 16 copies, upon which fluorescence did not increase and mCherry signals declined. This was due to the combined effects of RNA aptamer aggregation and reduced RNA stability. Unfortunately, both cellular and cytomegalovirus genomes were unable to maintain and express high Broccoli concatemer copy numbers, possibly due to recombination events. Interestingly, negative effects of Broccoli concatemers could be partially rescued by introducing linker sequences in between Broccoli repeats warranting further studies. Finally, we show that even though substrate-bound Broccoli is easily photobleached, it can still be utilized in live-cell imaging by adapting a time-lapse imaging protocol.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

