Cryo-EM of NHEJ supercomplexes provides insights into DNA repair
- PMID: 34352203
- PMCID: PMC9006396
- DOI: 10.1016/j.molcel.2021.07.005
Cryo-EM of NHEJ supercomplexes provides insights into DNA repair
Abstract
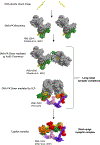
Non-homologous end joining (NHEJ) is one of two critical mechanisms utilized in humans to repair DNA double-strand breaks (DSBs). Unrepaired or incorrect repair of DSBs can lead to apoptosis or cancer. NHEJ involves several proteins, including the Ku70/80 heterodimer, DNA-dependent protein kinase catalytic subunit (DNA-PKcs), X-ray cross-complementing protein 4 (XRCC4), XRCC4-like factor (XLF), and ligase IV. These core proteins bind DSBs and ligate the damaged DNA ends. However, details of the structural assembly of these proteins remain unclear. Here, we present cryo-EM structures of NHEJ supercomplexes that are composed of these core proteins and DNA, revealing the detailed structural architecture of this assembly. We describe monomeric and dimeric forms of this supercomplex and also propose the existence of alternate dimeric forms of long-range synaptic complexes. Finally, we show that mutational disruption of several structural features within these NHEJ complexes negatively affects DNA repair.
Keywords: DNA repair; DNA-PK; NHEJ; cryo-EM; long-range synaptic complexes; non-homologous end joining.
Copyright © 2021 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests T.M.D.O. is employed at AstraZeneca.
Figures


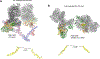


References
-
- Brouwer I, Sitters G, Candelli A, Heerema SJ, Heller I, de Melo AJ, Zhang H, Normanno D, Modesti M, Peterman EJ, and Wuite GJ (2016). Sliding sleeves of XRCC4-XLF bridge DNA and connect fragments of broken DNA. Nature 535, 566–569. - PubMed
-
- Bryans M, Valenzano MC, and Stamato TD (1999). Absence of DNA ligase IV protein in XR-1 cells: evidence for stabilization by XRCC4. Mutat. Res 433, 53–58. - PubMed
-
- Chaplin AK, Hardwick SW, Liang S, Kefala Stavridi A, Hnizda A, Cooper LR, De Oliveira TM, Chirgadze DY, and Blundell TL (2021). Dimers of DNA-PK create a stage for DNA double-strand break repair. Nat. Struct. Mol. Biol 28, 13–19. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

