At the Intersection of Major and Minor Spliceosomes: Crosstalk Mechanisms and Their Impact on Gene Expression
- PMID: 34354740
- PMCID: PMC8329584
- DOI: 10.3389/fgene.2021.700744
At the Intersection of Major and Minor Spliceosomes: Crosstalk Mechanisms and Their Impact on Gene Expression
Abstract
Many eukaryotic species contain two separate molecular machineries for removing non-coding intron sequences from pre-mRNA molecules. The majority of introns (more than 99.5% in humans) are recognized and excised by the major spliceosome, which utilizes relatively poorly conserved sequence elements at the 5' and 3' ends of the intron that are used for intron recognition and in subsequent catalysis. In contrast, the minor spliceosome targets a rare group of introns (approximately 0.5% in humans) with highly conserved sequences at the 5' and 3' ends of the intron. Minor introns coexist in the same genes with major introns and while the two intron types are spliced by separate spliceosomes, the two splicing machineries can interact with one another to shape mRNA processing events in genes containing minor introns. Here, we review known cooperative and competitive interactions between the two spliceosomes and discuss the mechanistic basis of the spliceosome crosstalk, its regulatory significance, and impact on spliceosome diseases.
Keywords: RNA processing; cryptic splicing; exon definition; mRNA splicing; major spliceosome; minor spliceosome; minor spliceosome disease.
Copyright © 2021 Akinyi and Frilander.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



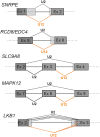
References
Publication types
LinkOut - more resources
Full Text Sources

