Investigation of Thermomorphogenesis-Related Genes for a Multi-Silique Trait in Brassica napus by Comparative Transcriptome Analysis
- PMID: 34367242
- PMCID: PMC8343136
- DOI: 10.3389/fgene.2021.678804
Investigation of Thermomorphogenesis-Related Genes for a Multi-Silique Trait in Brassica napus by Comparative Transcriptome Analysis
Abstract
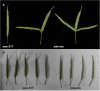
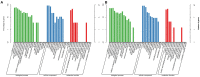
In higher plants, the structure of a flower is precisely controlled by a series of genes. An aberrance flower results in abnormal fruit morphology. Previously, we reported multi-silique rapeseed (Brassica napus) line zws-ms. We identified two associated regions and investigated differentially expressed genes (DEGs); thus, some candidate genes underlying the multi-silique phenotype in warm area Xindu were selected. However, this phenotype was switched off by lower temperature, and the responsive genes, known as thermomorphogenesis-related genes, remained elusive. So, based on that, in this study, we further investigated the transcriptome data from buds of zws-ms and its near-isogenic line zws-217 grown in colder area Ma'erkang, where both lines showed normal siliques only, and the DEGs between them analyzed. We compared the 129 DEGs from Xindu to the 117 ones from Ma'erkang and found that 33 of them represented the same or similar expression trends, whereas the other 96 DEGs showed different expression trends, which were defined as environment-specific. Furthermore, we combined this with the gene annotations and ortholog information and then selected BnaA09g45320D (chaperonin gene CPN10-homologous) and BnaC08g41780D [Seryl-tRNA synthetase gene OVULE ABORTION 7 (OVA7)-homologous] the possible thermomorphogenesis-related genes, which probably switched off the multi-silique under lower temperature. This study paves a way to a new perspective into flower/fruit development in Brassica plants.
Keywords: Brassica napus; RNA-seq; differentially expressed gene; environmental effect; multi-silique.
Copyright © 2021 Chai, Zhang, Li, Cui, Jiang, Zheng, Wu and Jiang.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Analysis of Altered Flowering Related Genes in a Multi-Silique Rapeseed (Brassica napus L.) Line zws-ms Based on Combination of Genome, Transcriptome and Proteome Data.Plants (Basel). 2023 Jun 23;12(13):2429. doi: 10.3390/plants12132429. Plants (Basel). 2023. PMID: 37446989 Free PMC article.
-
Investigation for a multi-silique trait in Brassica napus by alternative splicing analysis.PeerJ. 2020 Oct 8;8:e10135. doi: 10.7717/peerj.10135. eCollection 2020. PeerJ. 2020. PMID: 33083151 Free PMC article.
-
Identification of genomic regions associated with multi-silique trait in Brassica napus.BMC Genomics. 2019 Apr 23;20(1):304. doi: 10.1186/s12864-019-5675-4. BMC Genomics. 2019. PMID: 31014236 Free PMC article.
-
A systematic dissection of the mechanisms underlying the natural variation of silique number in rapeseed (Brassica napus L.) germplasm.Plant Biotechnol J. 2020 Feb;18(2):568-580. doi: 10.1111/pbi.13224. Epub 2019 Sep 17. Plant Biotechnol J. 2020. PMID: 31368615 Free PMC article.
-
Transcriptome analysis of Brassica napus pod using RNA-Seq and identification of lipid-related candidate genes.BMC Genomics. 2015 Oct 24;16:858. doi: 10.1186/s12864-015-2062-7. BMC Genomics. 2015. PMID: 26499887 Free PMC article.
Cited by
-
Analysis of Altered Flowering Related Genes in a Multi-Silique Rapeseed (Brassica napus L.) Line zws-ms Based on Combination of Genome, Transcriptome and Proteome Data.Plants (Basel). 2023 Jun 23;12(13):2429. doi: 10.3390/plants12132429. Plants (Basel). 2023. PMID: 37446989 Free PMC article.
-
Alternative Splicing Variation: Accessing and Exploiting in Crop Improvement Programs.Int J Mol Sci. 2023 Oct 15;24(20):15205. doi: 10.3390/ijms242015205. Int J Mol Sci. 2023. PMID: 37894886 Free PMC article. Review.
-
The Multi-Pistil Phenomenon in Higher Plants.Plants (Basel). 2025 Apr 4;14(7):1125. doi: 10.3390/plants14071125. Plants (Basel). 2025. PMID: 40219193 Free PMC article. Review.
References
-
- Benjamini Y., Hochberg Y. (1995). Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. Roy. Stat. Soc. B 57 289–300. 10.1111/j.2517-6161.1995.tb02031.x - DOI
-
- Chai L., Zhang J., Li H., Jiang J., Cui C., Zheng B., et al. (2020b). Dynamic Transcriptome Analysis of A Multi-Silique Trait in Rapeseed (Brassica napus L.). Int. J. Agric. Biol. 24 1625–1632.
LinkOut - more resources
Full Text Sources

