Comparative Analysis of Promoters and Enhancers in the Pituitary Glands of the Bama Xiang and Large White Pigs
- PMID: 34367256
- PMCID: PMC8343535
- DOI: 10.3389/fgene.2021.697994
Comparative Analysis of Promoters and Enhancers in the Pituitary Glands of the Bama Xiang and Large White Pigs
Abstract
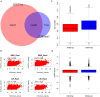
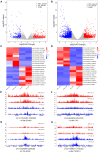
The epigenetic regulation of gene expression is implicated in complex diseases in humans and various phenotypes in other species. There has been little exploration of regulatory elements in the pig. Here, we performed chromatin immunoprecipitation coupled with high-throughput sequencing (ChIP-Seq) to profile histone H3 lysine 4 trimethylation (H3K4me3) and histone H3 lysine 27 acetylation (H3K27ac) in the pituitary gland of adult Bama Xiang and Large White pigs, which have divergent evolutionary histories and large phenotypic differences. We identified a total of 65,044 non-redundant regulatory regions, including 23,680 H3K4me3 peaks and 61,791 H3K27ac peaks (12,318 proximal and 49,473 distal), augmenting the catalog of pituitary regulatory elements in pigs. We found 793 H3K4me3 and 3,602 H3K27ac peaks that show differential activity between the two breeds, overlapping with genes involved in the Notch signaling pathway, response to growth hormone (GH), thyroid hormone signaling pathway, and immune system, and enriched for binding motifs of transcription factors (TFs), including JunB, ATF3, FRA1, and BATF. We further identified 2,025 non-redundant super enhancers from H3K27ac ChIP-seq data, among which 302 were shared in all samples of cover genes enriched for biological processes related to pituitary function. This study generated a valuable dataset of H3K4me3 and H3K27ac regions in porcine pituitary glands and revealed H3K4me3 and H3K27ac peaks with differential activity between Bama Xiang and Large White pigs.
Keywords: ChIP-Seq; H3K27ac; H3K4me3; differential peak activity; pigs; pituitary gland; super enhancer.
Copyright © 2021 Zhou, Zhu, Zhang, Jiang, Ling, Yang and Li.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
A comparative investigation on H3K27ac enhancer activities in the brain and liver tissues between wild boars and domesticated pigs.Evol Appl. 2022 Aug 16;15(8):1281-1290. doi: 10.1111/eva.13461. eCollection 2022 Aug. Evol Appl. 2022. PMID: 36051459 Free PMC article.
-
The super-enhancer repertoire in porcine liver.J Anim Sci. 2023 Jan 3;101:skad056. doi: 10.1093/jas/skad056. J Anim Sci. 2023. PMID: 36800318 Free PMC article.
-
The hyper-activation of transcriptional enhancers in breast cancer.Clin Epigenetics. 2019 Mar 12;11(1):48. doi: 10.1186/s13148-019-0645-x. Clin Epigenetics. 2019. PMID: 30867030 Free PMC article.
-
STAT3 Facilitates Super Enhancer Formation to Promote Fibroblast-To-Myofibroblast Differentiation by the Analysis of ATAC-Seq, RNA-Seq and ChIP-Seq.J Cell Mol Med. 2025 Jun;29(11):e70639. doi: 10.1111/jcmm.70639. J Cell Mol Med. 2025. PMID: 40464702 Free PMC article. Review.
-
Broad domains of histone H3 lysine 4 trimethylation in transcriptional regulation and disease.FEBS J. 2020 Jul;287(14):2891-2902. doi: 10.1111/febs.15219. Epub 2020 Feb 4. FEBS J. 2020. PMID: 31967712 Review.
Cited by
-
PorcineAI-Enhancer: Prediction of Pig Enhancer Sequences Using Convolutional Neural Networks.Animals (Basel). 2023 Sep 15;13(18):2935. doi: 10.3390/ani13182935. Animals (Basel). 2023. PMID: 37760334 Free PMC article.
-
Comparative Analyses of Sperm DNA Methylomes Among Three Commercial Pig Breeds Reveal Vital Hypomethylated Regions Associated With Spermatogenesis and Embryonic Development.Front Genet. 2021 Oct 6;12:740036. doi: 10.3389/fgene.2021.740036. eCollection 2021. Front Genet. 2021. PMID: 34691153 Free PMC article.
-
The landscape of super-enhancer regulates remote target gene transcription through loop domains in adipose tissue of pig.Heliyon. 2024 Feb 9;10(4):e25725. doi: 10.1016/j.heliyon.2024.e25725. eCollection 2024 Feb 29. Heliyon. 2024. PMID: 38390098 Free PMC article.
-
Tissue-Specific Chromatin Accessibility Regions and Transcription Factor Binding Sites in Pig Brain and Endocrine Tissues.Mol Neurobiol. 2025 Jun 7. doi: 10.1007/s12035-025-05128-5. Online ahead of print. Mol Neurobiol. 2025. PMID: 40483385
-
Macrophage-Centric Immunometabolic Crosstalk in Alopecia Areata Pathogenesis: Mechanisms and Therapeutic Implications.Clin Rev Allergy Immunol. 2025 May 22;68(1):50. doi: 10.1007/s12016-025-09060-3. Clin Rev Allergy Immunol. 2025. PMID: 40405030 Review.
References
-
- Ashery-Padan R., Alvarez-Bolado G., Klamt B., Gessler M., Gruss P. (1999). Fjx1, the murine homologue of the Drosophila four-jointed gene, codes for a putative secreted protein expressed in restricted domains of the developing and adult brain. Mech. Dev. 80 213–217. - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

