Chemical approaches to discover the full potential of peptide nucleic acids in biomedical applications
- PMID: 34367346
- PMCID: PMC8313981
- DOI: 10.3762/bjoc.17.116
Chemical approaches to discover the full potential of peptide nucleic acids in biomedical applications
Abstract
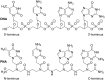
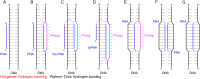
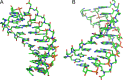
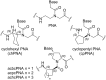
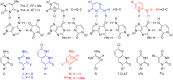
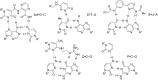
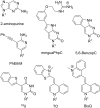
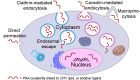
Peptide nucleic acid (PNA) is arguably one of the most successful DNA mimics, despite a most dramatic departure from the native structure of DNA. The present review summarizes 30 years of research on PNA's chemistry, optimization of structure and function, applications as probes and diagnostics, and attempts to develop new PNA therapeutics. The discussion starts with a brief review of PNA's binding modes and structural features, followed by the most impactful chemical modifications, PNA enabled assays and diagnostics, and discussion of the current state of development of PNA therapeutics. While many modifications have improved on PNA's binding affinity and specificity, solubility and other biophysical properties, the original PNA is still most frequently used in diagnostic and other in vitro applications. Development of therapeutics and other in vivo applications of PNA has notably lagged behind and is still limited by insufficient bioavailability and difficulties with tissue specific delivery. Relatively high doses are required to overcome poor cellular uptake and endosomal entrapment, which increases the risk of toxicity. These limitations remain unsolved problems waiting for innovative chemistry and biology to unlock the full potential of PNA in biomedical applications.
Keywords: PNA; antisense; chemical modifications; diagnostics; peptide nucleic acid.
Copyright © 2021, Brodyagin et al.
Figures



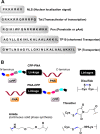
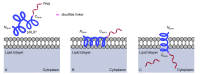

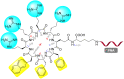
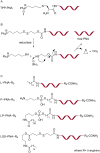



References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
