Evaluating CRISPR-based prime editing for cancer modeling and CFTR repair in organoids
- PMID: 34373320
- PMCID: PMC8356249
- DOI: 10.26508/lsa.202000940
Evaluating CRISPR-based prime editing for cancer modeling and CFTR repair in organoids
Abstract
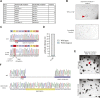
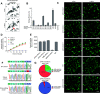
Prime editing is a recently reported genome editing tool using a nickase-cas9 fused to a reverse transcriptase that directly synthesizes the desired edit at the target site. Here, we explore the use of prime editing in human organoids. Common TP53 mutations can be correctly modeled in human adult stem cell-derived colonic organoids with efficiencies up to 25% and up to 97% in hepatocyte organoids. Next, we functionally repaired the cystic fibrosis CFTR-F508del mutation and compared prime editing to CRISPR/Cas9-mediated homology-directed repair and adenine base editing on the CFTR-R785* mutation. Whole-genome sequencing of prime editing-repaired organoids revealed no detectable off-target effects. Despite encountering varying editing efficiencies and undesired mutations at the target site, these results underline the broad applicability of prime editing for modeling oncogenic mutations and showcase the potential clinical application of this technique, pending further optimization.
© 2021 Geurts et al.
Conflict of interest statement
JM Beekman is an inventor on (a) patent(s) related to the FIS assay and received financial royalties from 2017 onward. JM Beekman reports receiving (a) research grant(s) and consultancy fees from various industries, including Vertex Pharmaceuticals, Proteostasis Therapeutics, Eloxx Pharmaceuticals, Teva Pharmaceutical Industries, and Galapagos outside the submitted work. H Clevers holds several patents on organoid technology. Their application numbers are as follows: PCT/NL2008/050543, WO2009/022907; PCT/NL2010/000017, WO2010/090513; PCT/IB2011/002167, WO2012/014076; PCT/IB2012/052950, WO2012/168930; PCT/EP2015/060815, WO2015/173425; PCT/EP2015/077990, WO2016/083613; PCT/EP2015/077988, WO2016/083612; PCT/EP2017/054797, WO2017/149025; PCT/EP2017/065101, WO2017/220586; PCT/EP2018/086716; and GB1819224.5.
Figures








References
-
- Aida T, Wilde JJ, Yang L, Hou Y, Li M, Xu D, Lin J, Qi P, Lu Z, Feng G (2020) Prime editing primarily induces undesired outcomes in mice. BioRxiv 10.1101/2020.08.06.239723(Preprint posted August 6, 2020). - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
