From a genome assembly to full regulatory network prediction: the case study of Rhodotorula toruloides putative Haa1-regulon
- PMID: 34376148
- PMCID: PMC8353774
- DOI: 10.1186/s12859-021-04312-3
From a genome assembly to full regulatory network prediction: the case study of Rhodotorula toruloides putative Haa1-regulon
Abstract
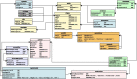
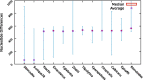
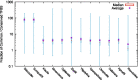
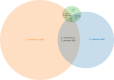
Numerous genomes are sequenced and made available to the community through the NCBI portal. However, and, unlike what happens for gene function annotation, annotation of promoter sequences and the underlying prediction of regulatory associations is mostly unavailable, severely limiting the ability to interpret genome sequences in a functional genomics perspective. Here we present an approach where one can download a genome of interest from NCBI in the GenBank Flat File (.gbff) format and, with a minimum set of commands, have all the information parsed, organized and made available through the platform web interface. Also, the new genomes are compared with a given genome of reference in search of homologous genes, shared regulatory elements and predicted transcription associations. We present this approach within the context of Community YEASTRACT of the YEASTRACT + portal, thus benefiting from immediate access to all the comparative genomics queries offered in the YEASTRACT + portal. Besides the yeast community, other communities can install the platform independently, without any constraints. In this work, we exemplify the usefulness of the presented tool, within Community YEASTRACT, in constructing a dedicated database and analysing the genome of the highly promising oleaginous red yeast species Rhodotorula toruloides currently poorly studied at the genome and transcriptome levels and with limited genome editing tools. Regulatory prediction is based on the conservation of promoter sequences and available regulatory networks. The case-study examined is focused on the Haa1 transcription factor-a key regulator of yeast resistance to acetic acid, an important inhibitor of industrial bioconversion of lignocellulosic hydrolysates. The new tool described here led to the prediction of a RtHaa1 regulon with expected impact in the optimization of R. toruloides robustness for lignocellulosic and pectin-rich residue biorefinery processes.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

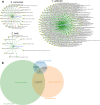
References
-
- Cherry JM, Hong EL, Amundsen C, Balakrishnan R, Binkley G, Chan ET, Christie KR, Costanzo MC, Dwight SS, Engel SR, Fisk DG, Hirschman JE, Hitz BC, Karra K, Krieger CJ, Miyasato SR, Nash RS, Park J, Skrzypek MS, Simison M, Weng S, Wong ED. Saccharomyces Genome Database: the genomics resource of budding yeast. Nucleic Acids Res. 2012;40:D700–D705. doi: 10.1093/nar/gkr1029. - DOI - PMC - PubMed
-
- Monteiro PT, Oliveira J, Pais P, Antunes M, Palma M, Cavalheiro M, Galocha M, Godinho CP, Martins LC, Bourbon N, Mota MN, Ribeiro RA, Viana R, Sá-Correia I, Teixeira MC. YEASTRACT+: a portal for cross-species comparative genomics of transcription regulation in yeasts. Nucleic Acids Res. 2020;48:D642–D649. doi: 10.1093/nar/gkz859. - DOI - PMC - PubMed
-
- High-density cultivation of oleaginous yeast Rhodosporidium toruloides Y4 in fed-batch culture. Enzyme Microb Technol. 2007;41:312–7.
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

