The transcription factor CmLEC1 positively regulates the seed-setting rate in hybridization breeding of chrysanthemum
- PMID: 34376645
- PMCID: PMC8355372
- DOI: 10.1038/s41438-021-00625-9
The transcription factor CmLEC1 positively regulates the seed-setting rate in hybridization breeding of chrysanthemum
Abstract
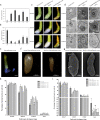
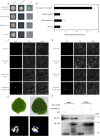
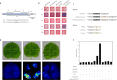
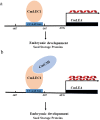
Distant hybridization is widely used to develop crop cultivars, whereas the hybridization process of embryo abortion often severely reduces the sought-after breeding effect. The LEAFY COTYLEDON1 (LEC1) gene has been extensively investigated as a central regulator of seed development, but it is far less studied in crop hybridization breeding. Here we investigated the function and regulation mechanism of CmLEC1 from Chrysanthemum morifolium during its seed development in chrysanthemum hybridization. CmLEC1 encodes a nucleic protein and is specifically expressed in embryos. CmLEC1's overexpression significantly promoted the seed-setting rate of the cross, while the rate was significantly decreased in the amiR-CmLEC1 transgenic chrysanthemum. The RNA-Seq analysis of the developing hybrid embryos revealed that regulatory genes involved in seed development, namely, CmLEA (late embryogenesis abundant protein), CmOLE (oleosin), CmSSP (seed storage protein), and CmEM (embryonic protein), were upregulated in the OE (overexpressing) lines but downregulated in the amiR lines vs. wild-type lines. Future analysis demonstrated that CmLEC1 directly activated CmLEA expression and interacted with CmC3H, and this CmLEC1-CmC3H interaction could enhance the transactivation ability of CmLEC1 for the expression of CmLEA. Further, CmLEC1 was able to induce several other key genes related to embryo development. Taken together, our results show that CmLEC1 plays a positive role in the hybrid embryo development of chrysanthemum plants, which might involve activating CmLEA's expression and interacting with CmC3H. This may be a new pathway in the LEC1 regulatory network to promote seed development, one perhaps leading to a novel strategy to not only overcome embryo abortion during crop breeding but also increase the seed yield.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Chrysanthemum embryo development is negatively affected by a novel ERF transcription factor, CmERF12.J Exp Bot. 2022 Jan 5;73(1):197-212. doi: 10.1093/jxb/erab398. J Exp Bot. 2022. PMID: 34453430
-
Chromosome doubling to overcome the chrysanthemum cross barrier based on insight from transcriptomic and proteomic analyses.BMC Genomics. 2016 Aug 9;17:585. doi: 10.1186/s12864-016-2939-0. BMC Genomics. 2016. PMID: 27506621 Free PMC article.
-
Identification of MicroRNAs and their Targets Associated with Embryo Abortion during Chrysanthemum Cross Breeding via High-Throughput Sequencing.PLoS One. 2015 Apr 24;10(4):e0124371. doi: 10.1371/journal.pone.0124371. eCollection 2015. PLoS One. 2015. PMID: 25909659 Free PMC article.
-
Anthocyanins in Floral Colors: Biosynthesis and Regulation in Chrysanthemum Flowers.Int J Mol Sci. 2020 Sep 7;21(18):6537. doi: 10.3390/ijms21186537. Int J Mol Sci. 2020. PMID: 32906764 Free PMC article. Review.
-
Regulation and evolution of the interaction of the seed B3 transcription factors with NF-Y subunits.Biochim Biophys Acta Gene Regul Mech. 2017 Oct;1860(10):1069-1078. doi: 10.1016/j.bbagrm.2017.08.008. Epub 2017 Sep 1. Biochim Biophys Acta Gene Regul Mech. 2017. PMID: 28866096 Review.
Cited by
-
A LlWRKY33-LlHSFA4-LlCAT2 module confers resistance to Botrytis cinerea in lily.Hortic Res. 2023 Nov 27;11(1):uhad254. doi: 10.1093/hr/uhad254. eCollection 2024 Jan. Hortic Res. 2023. PMID: 38274648 Free PMC article.
-
Low LdMYB12 expression contributes to petal spot deficiency in Lilium davidii var. unicolor.Mol Genet Genomics. 2023 Nov;298(6):1545-1557. doi: 10.1007/s00438-023-02080-8. Epub 2023 Nov 1. Mol Genet Genomics. 2023. PMID: 37910265
-
LoBLH6 interacts with LoMYB65 to regulate anther development through feedback regulation of gibberellin synthesis in lily.Hortic Res. 2024 Dec 4;12(3):uhae339. doi: 10.1093/hr/uhae339. eCollection 2025 Mar. Hortic Res. 2024. PMID: 40061809 Free PMC article.
-
RNA sequencing analysis reveals PgbHLH28 as the key regulator in response to methyl jasmonate-induced saponin accumulation in Platycodon grandiflorus.Hortic Res. 2024 Feb 28;11(5):uhae058. doi: 10.1093/hr/uhae058. eCollection 2024 May. Hortic Res. 2024. PMID: 38716227 Free PMC article.
References
-
- Kaul, M. L. H. Male Sterility in Higher Plants.Monographs on Theoretical and Applied Genetics (Springer, 1988).

