The sugar-responsive enteroendocrine neuropeptide F regulates lipid metabolism through glucagon-like and insulin-like hormones in Drosophila melanogaster
- PMID: 34376687
- PMCID: PMC8355161
- DOI: 10.1038/s41467-021-25146-w
The sugar-responsive enteroendocrine neuropeptide F regulates lipid metabolism through glucagon-like and insulin-like hormones in Drosophila melanogaster
Abstract
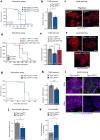
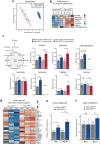
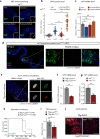
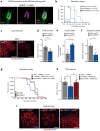
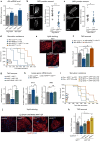
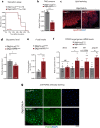
The enteroendocrine cell (EEC)-derived incretins play a pivotal role in regulating the secretion of glucagon and insulins in mammals. Although glucagon-like and insulin-like hormones have been found across animal phyla, incretin-like EEC-derived hormones have not yet been characterised in invertebrates. Here, we show that the midgut-derived hormone, neuropeptide F (NPF), acts as the sugar-responsive, incretin-like hormone in the fruit fly, Drosophila melanogaster. Secreted NPF is received by NPF receptor in the corpora cardiaca and in insulin-producing cells. NPF-NPFR signalling resulted in the suppression of the glucagon-like hormone production and the enhancement of the insulin-like peptide secretion, eventually promoting lipid anabolism. Similar to the loss of incretin function in mammals, loss of midgut NPF led to significant metabolic dysfunction, accompanied by lipodystrophy, hyperphagia, and hypoglycaemia. These results suggest that enteroendocrine hormones regulate sugar-dependent metabolism through glucagon-like and insulin-like hormones not only in mammals but also in insects.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
Gut-to-brain regulation of Drosophila aging through neuropeptide F, insulin, and juvenile hormone.Proc Natl Acad Sci U S A. 2024 Oct 22;121(43):e2411987121. doi: 10.1073/pnas.2411987121. Epub 2024 Oct 16. Proc Natl Acad Sci U S A. 2024. PMID: 39413128 Free PMC article.
-
Taste signaling elements expressed in gut enteroendocrine cells regulate nutrient-responsive secretion of gut hormones.Am J Clin Nutr. 2009 Sep;90(3):822S-825S. doi: 10.3945/ajcn.2009.27462T. Epub 2009 Jul 1. Am J Clin Nutr. 2009. PMID: 19571229 Free PMC article.
-
Midgut-derived neuropeptide F controls germline stem cell proliferation in a mating-dependent manner.PLoS Biol. 2018 Sep 24;16(9):e2005004. doi: 10.1371/journal.pbio.2005004. eCollection 2018 Sep. PLoS Biol. 2018. PMID: 30248087 Free PMC article.
-
Role of Incretin Hormones in Bowel Diseases.Endocr Dev. 2017;32:49-73. doi: 10.1159/000475731. Epub 2017 Aug 15. Endocr Dev. 2017. PMID: 28873384 Review.
-
Glucose-dependent insulinotropic polypeptide: effects on insulin and glucagon secretion in humans.Dan Med J. 2016 Apr;63(4):B5230. Dan Med J. 2016. PMID: 27034187 Review.
Cited by
-
Cholecystokinin/sulfakinin peptide signaling: conserved roles at the intersection between feeding, mating and aggression.Cell Mol Life Sci. 2022 Mar 14;79(3):188. doi: 10.1007/s00018-022-04214-4. Cell Mol Life Sci. 2022. PMID: 35286508 Free PMC article. Review.
-
Sex determination gene transformer regulates the male-female difference in Drosophila fat storage via the adipokinetic hormone pathway.Elife. 2021 Oct 21;10:e72350. doi: 10.7554/eLife.72350. Elife. 2021. PMID: 34672260 Free PMC article.
-
Metabolic control of enteroendocrine cell fate through a redox state sensor CtBP.bioRxiv [Preprint]. 2025 Jul 2:2025.06.30.662346. doi: 10.1101/2025.06.30.662346. bioRxiv. 2025. PMID: 40631244 Free PMC article. Preprint.
-
Interorgan communication through peripherally derived peptide hormones in Drosophila.Fly (Austin). 2022 Dec;16(1):152-176. doi: 10.1080/19336934.2022.2061834. Fly (Austin). 2022. PMID: 35499154 Free PMC article. Review.
-
Gut AstA mediates sleep deprivation-induced energy wasting in Drosophila.Cell Discov. 2023 May 23;9(1):49. doi: 10.1038/s41421-023-00541-3. Cell Discov. 2023. PMID: 37221172 Free PMC article.
References
-
- Bansal P, Wang Q. Insulin as a physiological modulator of glucagon secretion. Am. J. Physiol. Metab. 2008;295:E751–E761. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

