Genetic and phenotypic analysis of the pathogenic potential of two novel Chlamydia gallinacea strains compared to Chlamydia psittaci
- PMID: 34389764
- PMCID: PMC8363750
- DOI: 10.1038/s41598-021-95966-9
Genetic and phenotypic analysis of the pathogenic potential of two novel Chlamydia gallinacea strains compared to Chlamydia psittaci
Abstract
Chlamydia gallinacea is an obligate intracellular bacterium that has recently been added to the family of Chlamydiaceae. C. gallinacea is genetically diverse, widespread in poultry and a suspected cause of pneumonia in slaughterhouse workers. In poultry, C. gallinacea infections appear asymptomatic, but studies about the pathogenic potential are limited. In this study two novel sequence types of C. gallinacea were isolated from apparently healthy chickens. Both isolates (NL_G47 and NL_F725) were closely related to each other and have at least 99.5% DNA sequence identity to C. gallinacea Type strain 08-1274/3. To gain further insight into the pathogenic potential, infection experiments in embryonated chicken eggs and comparative genomics with Chlamydia psittaci were performed. C. psittaci is a ubiquitous zoonotic pathogen of birds and mammals, and infection in poultry can result in severe systemic illness. In experiments with embryonated chicken eggs, C. gallinacea induced mortality was observed, potentially strain dependent, but lower compared to C. psittaci induced mortality. Comparative analyses confirmed all currently available C. gallinacea genomes possess the hallmark genes coding for known and potential virulence factors as found in C. psittaci albeit to a reduced number of orthologues or paralogs. The presence of potential virulence factors and the observed mortality in embryonated eggs indicates C. gallinacea should rather be considered as an opportunistic pathogen than an innocuous commensal.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
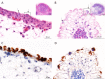



References
-
- Sachse K, Laroucau K, Vanrompay D. Avian chlamydiosis. Curr. Clin. Microbiol. Rep. 2015;2:10–21. doi: 10.1007/s40588-014-0010-y. - DOI
Publication types
MeSH terms
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

