Epigenetic Reprogramming in Early Animal Development
- PMID: 34400552
- PMCID: PMC9248830
- DOI: 10.1101/cshperspect.a039677
Epigenetic Reprogramming in Early Animal Development
Abstract
Dramatic nuclear reorganization occurs during early development to convert terminally differentiated gametes to a totipotent zygote, which then gives rise to an embryo. Aberrant epigenome resetting severely impairs embryo development and even leads to lethality. How the epigenomes are inherited, reprogrammed, and reestablished in this critical developmental period has gradually been unveiled through the rapid development of technologies including ultrasensitive chromatin analysis methods. In this review, we summarize the latest findings on epigenetic reprogramming in gametogenesis and embryogenesis, and how it contributes to gamete maturation and parental-to-zygotic transition. Finally, we highlight the key questions that remain to be answered to fully understand chromatin regulation and nuclear reprogramming in early development.
Copyright © 2022 Cold Spring Harbor Laboratory Press; all rights reserved.
Figures

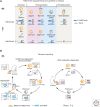
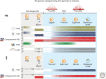
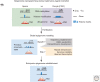
References
-
- Aguirre-Lavin T, Adenot P, Bonnet-Garnier A, Lehmann G, Fleurot R, Boulesteix C, Debey P, Beaujean N. 2012. 3D-FISH analysis of embryonic nuclei in mouse highlights several abrupt changes of nuclear organization during preimplantation development. BMC Dev Biol 12: 30. 10.1186/1471-213X-12-30 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
