GRIK2 is a target for bladder cancer stem-like cell-targeting immunotherapy
- PMID: 34405274
- PMCID: PMC10992913
- DOI: 10.1007/s00262-021-03025-z
GRIK2 is a target for bladder cancer stem-like cell-targeting immunotherapy
Abstract
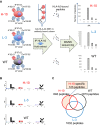
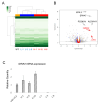
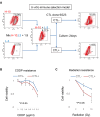
Recent studies have revealed that treatment-resistant cancer stem-like cells (CSCs)/cancer-initiating cells (CICs) can be targeted by cytotoxic T lymphocytes (CTLs). CTLs recognize antigenic peptides derived from tumor-associated antigens; thus, the identification of tumor-associated antigens expressed by CSCs/CICs is essential. Human leucocyte antigen (HLA) ligandome analysis using mass spectrometry enables the analysis of naturally expressed antigenic peptides; however, HLA ligandome analysis requires a large number of cells and is challenging for CSCs/CICs. In this study, we established a novel bladder CSC/CIC model from a bladder cancer cell line (UM-UC-3 cells) using an ALDEFLUOR assay. CSCs/CICs were isolated as aldehyde dehydrogenase (ALDH)-high cells and several ALDHhigh clone cells were established. ALDHhigh clone cells were enriched with CSCs/CICs by sphere formation and tumorigenicity in immunodeficient mice. HLA ligandome analysis and cap analysis of gene expression using ALDHhigh clone cells revealed a distinctive antigenic peptide repertoire in bladder CSCs/CICs, and we found that a glutamate receptor, ionotropic, kainite 2 (GRIK2)-derived antigenic peptide (LMYDAVHVV) was specifically expressed by CSCs/CICs. A GRIK2 peptide-specific CTL clone recognized GRIK2-overexpressing UM-UC-3 cells and ALDHhigh clone cells, indicating that GRIK2 peptide can be a novel target for bladder CSC/CIC-targeting immunotherapy.
Keywords: Antigen; Bladder cancer; Cancer stem cell; Cytotoxic T lymphocytes; GRIK2.
© 2021. The Author(s), under exclusive licence to Springer-Verlag GmbH Germany, part of Springer Nature.
Conflict of interest statement
The authors have no financial conflicts of interest to disclose.
Figures


Similar articles
-
Aldehyde Dehydrogenese-1 High Cancer Stem-like Cells/Cancer-initiating Cells Escape from Cytotoxic T Lymphocytes due to Lower Expression of Human Leukocyte Antigen Class 1.Anticancer Res. 2024 May;44(5):1877-1883. doi: 10.21873/anticanres.16989. Anticancer Res. 2024. PMID: 38677758
-
GRIK2 has a role in the maintenance of urothelial carcinoma stem-like cells, and its expression is associated with poorer prognosis.Oncotarget. 2017 Apr 25;8(17):28826-28839. doi: 10.18632/oncotarget.16259. Oncotarget. 2017. PMID: 28418868 Free PMC article.
-
Brother of the regulator of the imprinted site (BORIS) variant subfamily 6 is involved in cervical cancer stemness and can be a target of immunotherapy.Oncotarget. 2016 Mar 8;7(10):11223-37. doi: 10.18632/oncotarget.7165. Oncotarget. 2016. PMID: 26849232 Free PMC article.
-
Immunomodulating and Immunoresistance Properties of Cancer-Initiating Cells: Implications for the Clinical Success of Immunotherapy.Immunol Invest. 2017 Apr;46(3):221-238. doi: 10.1080/08820139.2017.1280051. Epub 2017 Mar 13. Immunol Invest. 2017. PMID: 28287848 Review.
-
Concise Review: Targeting Cancer Stem Cells Using Immunologic Approaches.Stem Cells. 2015 Jul;33(7):2085-92. doi: 10.1002/stem.2039. Epub 2015 May 13. Stem Cells. 2015. PMID: 25873269 Free PMC article. Review.
Cited by
-
Membrane-associated RING-CH 7 inhibits stem-like capacities of bladder cancer cells by interacting with nucleotide-binding oligomerization domain containing 1.Cell Biosci. 2024 Mar 10;14(1):32. doi: 10.1186/s13578-024-01210-y. Cell Biosci. 2024. PMID: 38462600 Free PMC article.
-
MHC class I trafficking signal improves induction of cytotoxic T lymphocyte using artificial antigen presenting cells.Biochem Biophys Rep. 2025 Feb 19;41:101946. doi: 10.1016/j.bbrep.2025.101946. eCollection 2025 Mar. Biochem Biophys Rep. 2025. PMID: 40046256 Free PMC article.
-
Establishment of potent TCR-T cells specific for cisplatin-resistance related tumor-associated antigen, CLSPN using codon-optimization.Hum Vaccin Immunother. 2024 Dec 31;20(1):2414542. doi: 10.1080/21645515.2024.2414542. Epub 2024 Nov 13. Hum Vaccin Immunother. 2024. PMID: 39539024 Free PMC article.
-
Restoration of ARID1A Protein in ARID1A-deficient Clear Cell Carcinoma of the Ovary Attenuates Reactivity to Cytotoxic T Lymphocytes.Cancer Genomics Proteomics. 2024 Jul-Aug;21(4):414-420. doi: 10.21873/cgp.20460. Cancer Genomics Proteomics. 2024. PMID: 38944423 Free PMC article.
-
Recent advances in bladder cancer stem cells (BCSCs): A descriptive review of emerging therapeutic targets.iScience. 2025 May 22;28(7):112720. doi: 10.1016/j.isci.2025.112720. eCollection 2025 Jul 18. iScience. 2025. PMID: 40635786 Free PMC article. Review.
References
-
- Advanced Bladder Cancer (ABC) Meta-analysis Collaboration (2005) Adjuvant chemotherapy in invasive bladder cancer: a systematic review and meta-analysis of individual patient data Advanced Bladder Cancer (ABC) Meta-analysis Collaboration. Eur Urol 48:189–199. Discussion 199–201 - PubMed
-
- International Collaboration of Trialists, Medical Research Council Advanced Bladder Cancer Working Party (now the National Cancer Research Institute Bladder Cancer Clinical Studies Group), European Organisation for Research and Treatment of Cancer Genito-Urinary Tract Cancer Group et al. (2011) International phase III trial assessing neoadjuvant cisplatin, methotrexate, and vinblastine chemotherapy for muscle-invasive bladder cancer: long-term results of the BA06 30894 trial. J Clin Oncol 29:2171–2177 - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

