Sex dependent glial-specific changes in the chromatin accessibility landscape in late-onset Alzheimer's disease brains
- PMID: 34429139
- PMCID: PMC8383438
- DOI: 10.1186/s13024-021-00481-0
Sex dependent glial-specific changes in the chromatin accessibility landscape in late-onset Alzheimer's disease brains
Abstract
Background: In the post-GWAS era, there is an unmet need to decode the underpinning genetic etiologies of late-onset Alzheimer's disease (LOAD) and translate the associations to causation.
Methods: We conducted ATAC-seq profiling using NeuN sorted-nuclei from 40 frozen brain tissues to determine LOAD-specific changes in chromatin accessibility landscape in a cell-type specific manner.
Results: We identified 211 LOAD-specific differential chromatin accessibility sites in neuronal-nuclei, four of which overlapped with LOAD-GWAS regions (±100 kb of SNP). While the non-neuronal nuclei did not show LOAD-specific differences, stratification by sex identified 842 LOAD-specific chromatin accessibility sites in females. Seven of these sex-dependent sites in the non-neuronal samples overlapped LOAD-GWAS regions including APOE. LOAD loci were functionally validated using single-nuclei RNA-seq datasets.
Conclusions: Using brain sorted-nuclei enabled the identification of sex-dependent cell type-specific LOAD alterations in chromatin structure. These findings enhance the interpretation of LOAD-GWAS discoveries, provide potential pathomechanisms, and suggest novel LOAD-loci.
Keywords: ATAC-seq; Chromatin accessibility; Gene dysregulation; Late-onset Alzheimer’s disease; Nuclei sorting; snRNA-seq.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
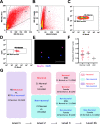




References
-
- Harold D, Abraham R, Hollingworth P, Sims R, Gerrish A, Hamshere ML, Pahwa JS, Moskvina V, Dowzell K, Williams A, Jones N, Thomas C, Stretton A, Morgan AR, Lovestone S, Powell J, Proitsi P, Lupton MK, Brayne C, Rubinsztein DC, Gill M, Lawlor B, Lynch A, Morgan K, Brown KS, Passmore PA, Craig D, McGuinness B, Todd S, Holmes C, Mann D, Smith AD, Love S, Kehoe PG, Hardy J, Mead S, Fox N, Rossor M, Collinge J, Maier W, Jessen F, Schürmann B, Heun R, van den Bussche H, Heuser I, Kornhuber J, Wiltfang J, Dichgans M, Frölich L, Hampel H, Hüll M, Rujescu D, Goate AM, Kauwe JSK, Cruchaga C, Nowotny P, Morris JC, Mayo K, Sleegers K, Bettens K, Engelborghs S, de Deyn PP, van Broeckhoven C, Livingston G, Bass NJ, Gurling H, McQuillin A, Gwilliam R, Deloukas P, al-Chalabi A, Shaw CE, Tsolaki M, Singleton AB, Guerreiro R, Mühleisen TW, Nöthen MM, Moebus S, Jöckel KH, Klopp N, Wichmann HE, Carrasquillo MM, Pankratz VS, Younkin SG, Holmans PA, O'Donovan M, Owen MJ, Williams J. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer's disease. Nat Genet. 2009;41(10):1088–1093. doi: 10.1038/ng.440. - DOI - PMC - PubMed
-
- Jansen I, Savage J, Watanabe K, Bryois J, Williams D, Steinberg S, et al. Genetic meta-analysis identifies 9 novel loci and functional pathways for Alzheimers disease risk. bioRxiv. 2018:258533. 10.1101/258533.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

