Transcription Factors Interact with ABA through Gene Expression and Signaling Pathways to Mitigate Drought and Salinity Stress
- PMID: 34439825
- PMCID: PMC8393639
- DOI: 10.3390/biom11081159
Transcription Factors Interact with ABA through Gene Expression and Signaling Pathways to Mitigate Drought and Salinity Stress
Abstract
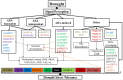
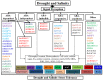
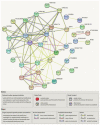
Among abiotic stressors, drought and salinity seriously affect crop growth worldwide. In plants, research has aimed to increase stress-responsive protein synthesis upstream or downstream of the various transcription factors (TFs) that alleviate drought and salinity stress. TFs play diverse roles in controlling gene expression in plants, which is necessary to regulate biological processes, such as development and environmental stress responses. In general, plant responses to different stress conditions may be either abscisic acid (ABA)-dependent or ABA-independent. A detailed understanding of how TF pathways and ABA interact to cause stress responses is essential to improve tolerance to drought and salinity stress. Despite previous progress, more active approaches based on TFs are the current focus. Therefore, the present review emphasizes the recent advancements in complex cascades of gene expression during drought and salinity responses, especially identifying the specificity and crosstalk in ABA-dependent and -independent signaling pathways. This review also highlights the transcriptional regulation of gene expression governed by various key TF pathways, including AP2/ERF, bHLH, bZIP, DREB, GATA, HD-Zip, Homeo-box, MADS-box, MYB, NAC, Tri-helix, WHIRLY, WOX, WRKY, YABBY, and zinc finger, operating in ABA-dependent and -independent signaling pathways.
Keywords: ABA; drought; genetic engineering; pathways; salinity; transcription factors.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

