Proteomic analysis of machine perfusion solution from brain dead donor kidneys reveals that elevated complement, cytoskeleton and lipid metabolism proteins are associated with 1-year outcome
- PMID: 34448265
- PMCID: PMC9292651
- DOI: 10.1111/tri.13984
Proteomic analysis of machine perfusion solution from brain dead donor kidneys reveals that elevated complement, cytoskeleton and lipid metabolism proteins are associated with 1-year outcome
Abstract
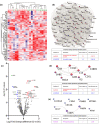
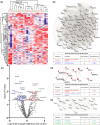
Assessment of donor kidney quality is based on clinical scores or requires biopsies for histological assessment. Noninvasive strategies to identify and predict graft outcome at an early stage are, therefore, needed. We evaluated the perfusate of donation after brain death (DBD) kidneys during nonoxygenated hypothermic machine perfusion (HMP). In particular, we compared perfusate protein profiles of good outcome (GO) and suboptimal outcome (SO) 1-year post-transplantation. Samples taken 15 min after the start HMP (T1) and before the termination of HMP (T2) were analysed using quantitative liquid chromatography-tandem mass spectrometry (LC-MS/MS). Hierarchical clustering of the 100 most abundant proteins showed discrimination between grafts with a GO and SO at T1. Elevated levels of proteins involved in classical complement cascades at both T1 and T2 and a reduced abundance of lipid metabolism at T1 and of cytoskeletal proteins at T2 in GO versus SO was observed. ATP-citrate synthase and fatty acid-binding protein 5 (T1) and immunoglobulin heavy variable 2-26 and desmoplakin (T2) showed 91% and 86% predictive values, respectively, for transplant outcome. Taken together, DBD kidney HMP perfusate profiles can distinguish between outcome 1-year post-transplantation. Furthermore, it provides insights into mechanisms that could play a role in post-transplant outcomes.
Keywords: biomarker discovery; complement activation; donation after brain death; hypothermic machine perfusion; kidney preservation; proteomics.
© 2021 The Authors. Transplant International published by John Wiley & Sons Ltd on behalf of Steunstichting ESOT.
Conflict of interest statement
The authors have declared no conflicts of interest.
Figures


References
-
- Eurotransplant . Eurotransplant annual report 2019. Leiden: 2019.
-
- Bos EM, Leuvenink HGD, van Goor H, Ploeg RJ. Kidney grafts from brain dead donors: Inferior quality or opportunity for improvement? Kidney Int 2007; 72(7): 797. - PubMed
-
- [4]Chen EP, Bittner HB, Kendall SW, Van Trigt P . Hormonal and hemodynamic changes in a validated animal model of brain death. Crit Care Med 1996; 24: 1352. - PubMed
-
- [5]Gavriilidis P, Inston NG. Recipient and allograft survival following donation after circulatory death versus donation after brain death for renal transplantation: a systematic review and meta‐analysis. Transpl Rev 2020; 34: 100563. - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

