A Tri-Stage Wrapper-Filter Feature Selection Framework for Disease Classification
- PMID: 34451013
- PMCID: PMC8402295
- DOI: 10.3390/s21165571
A Tri-Stage Wrapper-Filter Feature Selection Framework for Disease Classification
Abstract
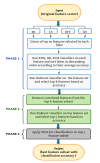
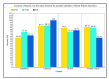
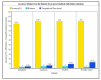
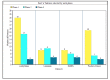
In machine learning and data science, feature selection is considered as a crucial step of data preprocessing. When we directly apply the raw data for classification or clustering purposes, sometimes we observe that the learning algorithms do not perform well. One possible reason for this is the presence of redundant, noisy, and non-informative features or attributes in the datasets. Hence, feature selection methods are used to identify the subset of relevant features that can maximize the model performance. Moreover, due to reduction in feature dimension, both training time and storage required by the model can be reduced as well. In this paper, we present a tri-stage wrapper-filter-based feature selection framework for the purpose of medical report-based disease detection. In the first stage, an ensemble was formed by four filter methods-Mutual Information, ReliefF, Chi Square, and Xvariance-and then each feature from the union set was assessed by three classification algorithms-support vector machine, naïve Bayes, and k-nearest neighbors-and an average accuracy was calculated. The features with higher accuracy were selected to obtain a preliminary subset of optimal features. In the second stage, Pearson correlation was used to discard highly correlated features. In these two stages, XGBoost classification algorithm was applied to obtain the most contributing features that, in turn, provide the best optimal subset. Then, in the final stage, we fed the obtained feature subset to a meta-heuristic algorithm, called whale optimization algorithm, in order to further reduce the feature set and to achieve higher accuracy. We evaluated the proposed feature selection framework on four publicly available disease datasets taken from the UCI machine learning repository, namely, arrhythmia, leukemia, DLBCL, and prostate cancer. Our obtained results confirm that the proposed method can perform better than many state-of-the-art methods and can detect important features as well. Less features ensure less medical tests for correct diagnosis, thus saving both time and cost.
Keywords: arrhythmia; cancer dataset; disease classification; feature selection; filter method; whale optimization algorithm; wrapper method.
Conflict of interest statement
The authors declare no conflict of interests.
Figures
References
-
- Ghosh M., Guha R., Singh P.K., Bhateja V., Sarkar R. A histogram based fuzzy ensemble technique for feature selection. Evol. Intell. 2019;12:713–724. doi: 10.1007/s12065-019-00279-6. - DOI
-
- Ghosh K.K., Ahmed S., Singh P.K., Geem Z.W., Sarkar R. Improved Binary Sailfish Optimizer Based on Adaptive β-Hill Climbing for Feature Selection. IEEE Access. 2020;8:83548–83560. doi: 10.1109/ACCESS.2020.2991543. - DOI
-
- Duval B., Hao J.-K., Hernandez J.C.H. A memetic algorithm for gene selection and molecular classification of cancer; Proceedings of the 11th Annual Conference on Genetic and Evolutionary Computation, GECCO ‘09; Montreal, QC, Canada. 8–12 July 2009; pp. 201–208. - DOI
-
- Chandrashekar G., Sahin F. A survey on feature selection methods. Comput. Electr. Eng. 2014;40:16–28. doi: 10.1016/j.compeleceng.2013.11.024. - DOI
-
- Lu H., Chen J., Yan K., Jin Q., Xue Y., Gao Z. A hybrid feature selection algorithm for gene expression data classification. Neurocomputing. 2017;256:56–62. doi: 10.1016/j.neucom.2016.07.080. - DOI
MeSH terms
LinkOut - more resources
Full Text Sources
Other Literature Sources

