Quantitative analyses for effects of neddylation on CRL2VHL substrate ubiquitination and degradation
- PMID: 34459035
- PMCID: PMC8521307
- DOI: 10.1002/pro.4176
Quantitative analyses for effects of neddylation on CRL2VHL substrate ubiquitination and degradation
Abstract
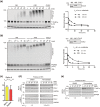
Through catalyzing the ubiquitination of key regulatory proteins, cullin-RING ubiquitin ligases (CRLs) play essential biological roles and their activities are controlled by multiple mechanisms including neddylation, the conjugation of NEDD8 to cullins. Upon neddylation, a CRL, such as the CUL1-based CRL1, undergoes conformational changes that accelerate substrate ubiquitination. Given the structural diversity across subfamilies of CRLs and their substrates, to what extent neddylation modulates the activity of individual CRLs remains to be evaluated. Here, through reconstituting the CRL2 ubiquitination reaction in vitro, we showed that neddylation promotes CRL2VHL -dependent degradation of both full-length HIF1α and the degron peptide of HIF1α, resulting in more than 10-fold increase in the rate of substrate ubiquitination. Consistently, pevonedistat (also known as MLN4924), an inhibitor of neddylation, inhibits the degradation of HIF1α in RCC4 cells stably expressing VHL in cycloheximide chase assays. However, such inhibitory effect of pevonedistat on HIF1α degradation was not observed in HEK293 cells, which was further found to be due to CRL2VHL -independent degradation that was active in HEK293 but not RCC4 cells. After truncating HIF1α to its Carboxy-terminal Oxygen-Dependent Degradation (CODD) domain, we showed that pevonedistat inhibited the degradation of CODD and increased its half-life by six-fold in HEK293 cells. Our results demonstrate that neddylation plays a significant role in activating CRL2, and the cellular activity of CRL2VHL is better reflected by the degradation of CODD than that of HIF1α, especially under conditions where CRL2-independent degradation of HIF1α is active.
Keywords: CODD; CRL2 ubiquitin ligase; neddylation; protein degradation; ubiquitination.
© 2021 The Authors. Protein Science published by Wiley Periodicals LLC on behalf of The Protein Society.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
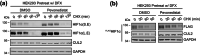

References
-
- Hershko A. The ubiquitin system for protein degradation. Annu Rev Biochem. 1992;61:761–807. - PubMed
-
- Varshavsky A. The ubiquitin system. Trends Biochem Sci. 1997;22:383–387. - PubMed
-
- Petroski MD, Deshaies RJ. Function and regulation of cullin‐RING ubiquitin ligases. Nat Rev Mol Cell Biol. 2005;6:9–20. - PubMed
-
- Zheng N, Schulman BA, Song L, et al. Structure of the Cul1‐Rbx1‐Skp1‐F boxSkp2 SCF ubiquitin ligase complex. Nature. 2002;416:703–709. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

