Whole-genome sequencing of endangered Zhoushan cattle suggests its origin and the association of MC1R with black coat colour
- PMID: 34462508
- PMCID: PMC8405626
- DOI: 10.1038/s41598-021-96896-2
Whole-genome sequencing of endangered Zhoushan cattle suggests its origin and the association of MC1R with black coat colour
Abstract
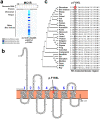
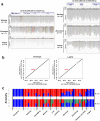
Zhoushan cattle are an endangered cattle breed in the Zhoushan islands in China. Since Zhoushan cattle have been bred in isolation, they show unique characteristics, such as dark black coat colour. However, no studies have been conducted on the genome of Zhoushan cattle. Here, we performed whole-genome sequencing of seven individuals of Zhoushan cattle and nine cattle in Wenling, geographically close to the Zhoushan islands. By integrating our data and publicly-available data, we found that Zhoushan cattle are genetically highly similar to Bos indicus cattle in south-eastern China. Furthermore, by identifying the genomic regions shared between Zhoushan cattle and Angus cattle, a Bos taurus breed, we found that the p.F195L mutation in melanocyte-stimulating hormone receptor (MC1R) could be associated with their dark black coat colour. Taken together, our results provide a valuable resource for characterising the uniqueness of Zhoushan cattle.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



References
-
- Porter V, Alderson L, Hall S, Sponenberg P. Mason’s World Encyclopedia of Livestock Breeds and Breeding. CAB International; 2016.
-
- Youlin J. Report on Domestic Animal Genetic Resources in China. China Agriculture Publishers; 2004.
-
- Jiang, Y. Where is the Chinese yellow cattle from?. Nat. Ecol. Evol.https://natureecoevocommunity.nature.com/posts/34309-where-is-the-chines.... (2018).
-
- Chen Y-C. Four interesting endangered breeds of animals in China. Anim. Genet. Resour. Inf. 1995;16:29–35. doi: 10.1017/S101423390000047X. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

