Environmental fluctuations reshape an unexpected diversity-disturbance relationship in a microbial community
- PMID: 34477107
- PMCID: PMC8460265
- DOI: 10.7554/eLife.67175
Environmental fluctuations reshape an unexpected diversity-disturbance relationship in a microbial community
Abstract
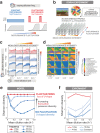
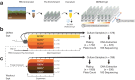
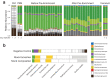
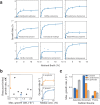
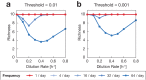
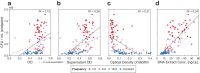
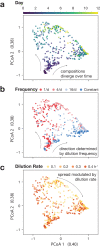
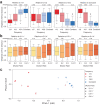
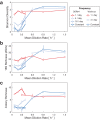
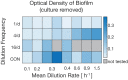
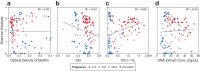
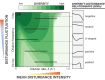
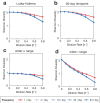
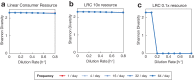
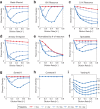
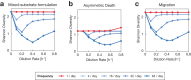
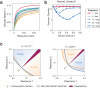
Environmental disturbances have long been theorized to play a significant role in shaping the diversity and composition of ecosystems. However, an inability to specify the characteristics of a disturbance experimentally has produced an inconsistent picture of diversity-disturbance relationships (DDRs). Here, using a high-throughput programmable culture system, we subjected a soil-derived bacterial community to dilution disturbance profiles with different intensities (mean dilution rates), applied either constantly or with fluctuations of different frequencies. We observed an unexpected U-shaped relationship between community diversity and disturbance intensity in the absence of fluctuations. Adding fluctuations increased community diversity and erased the U-shape. All our results are well-captured by a Monod consumer resource model, which also explains how U-shaped DDRs emerge via a novel 'niche flip' mechanism. Broadly, our combined experimental and modeling framework demonstrates how distinct features of an environmental disturbance can interact in complex ways to govern ecosystem assembly and offers strategies for reshaping the composition of microbiomes.
Keywords: bacterial communities; computational biology; continuous culture; microbial ecology; systems biology; systems modeling.
© 2021, Mancuso et al.
Conflict of interest statement
CM, HL, CA, JG No competing interests declared, AK is co-founder of Fynch Biosciences, a manufacturer of eVOLVER hardware.
Figures





References
-
- Apprill A, McNally S, Parsons R, Weber L. Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquatic Microbial Ecology. 2015;75:129–137. doi: 10.3354/ame01753. - DOI
-
- Bolyen E, Rideout JR, Dillon MR, Bokulich NA, Abnet CC, Al-Ghalith GA, Alexander H, Alm EJ, Arumugam M, Asnicar F, Bai Y, Bisanz JE, Bittinger K, Brejnrod A, Brislawn CJ, Brown CT, Callahan BJ, Caraballo-Rodríguez AM, Chase J, Cope EK, Da Silva R, Diener C, Dorrestein PC, Douglas GM, Durall DM, Duvallet C, Edwardson CF, Ernst M, Estaki M, Fouquier J, Gauglitz JM, Gibbons SM, Gibson DL, Gonzalez A, Gorlick K, Guo J, Hillmann B, Holmes S, Holste H, Huttenhower C, Huttley GA, Janssen S, Jarmusch AK, Jiang L, Kaehler BD, Kang KB, Keefe CR, Keim P, Kelley ST, Knights D, Koester I, Kosciolek T, Kreps J, Langille MGI, Lee J, Ley R, Liu YX, Loftfield E, Lozupone C, Maher M, Marotz C, Martin BD, McDonald D, McIver LJ, Melnik AV, Metcalf JL, Morgan SC, Morton JT, Naimey AT, Navas-Molina JA, Nothias LF, Orchanian SB, Pearson T, Peoples SL, Petras D, Preuss ML, Pruesse E, Rasmussen LB, Rivers A, Robeson MS, Rosenthal P, Segata N, Shaffer M, Shiffer A, Sinha R, Song SJ, Spear JR, Swafford AD, Thompson LR, Torres PJ, Trinh P, Tripathi A, Turnbaugh PJ, Ul-Hasan S, van der Hooft JJJ, Vargas F, Vázquez-Baeza Y, Vogtmann E, von Hippel M, Walters W, Wan Y, Wang M, Warren J, Weber KC, Williamson CHD, Willis AD, Xu ZZ, Zaneveld JR, Zhang Y, Zhu Q, Knight R, Caporaso JG. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature Biotechnology. 2019;37:852–857. doi: 10.1038/s41587-019-0209-9. - DOI - PMC - PubMed
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

