Characterization of a Novel Hepatitis C Subtype, 6xj, and Its Consequences for Direct-Acting Antiviral Treatment in Yunnan, China
- PMID: 34479413
- PMCID: PMC8552672
- DOI: 10.1128/Spectrum.00297-21
Characterization of a Novel Hepatitis C Subtype, 6xj, and Its Consequences for Direct-Acting Antiviral Treatment in Yunnan, China
Abstract
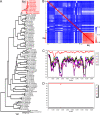
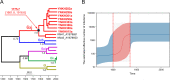
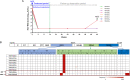
Hepatitis C virus (HCV) has a high rate of genetic variability, with eight genotypes and 91 subtypes. The genetic diversity of HCV genotype 6 (HCV-6) is the highest with 31 subtypes, and this genotype is prevalent in Southeast Asia. In this study, we investigated 160 individuals with chronic hepatitis C in Yunnan Province, China. Using reverse transcription (RT)-PCR and Sanger sequencing, 147 cases were successfully amplified and genotyped as 3b (4.9%), 3a (19.73%), 6n (12.24%), 1b (7.48%), 2a (6.12%), 6a (2.04%), 1a (0.68%), 6v (0.68%), and 6xa (0.68%), with eight sequences remaining unclassified. Subsequently, the eight nearly full-length genomes were successfully amplified and analyzed. The eight complete coding sequences formed a phylogenetic group that was distinct from the previously assigned HCV-6 subtypes and clustered with two previously unnamed HCV-6 sequences. Furthermore, Simplot analysis showed no recombination and the p-distance was more than 15% in comparison to the 6a to 6xi subtypes. Taken together, we identified a new HCV-6 subtype, 6xj, which originated approximately in 1775 according to Bayesian analyses. Moreover, all eight individuals received follow-up assessments at 44 weeks from the beginning of their 12-week treatments of sofosbuvir/velpatasvir (after-treatment week 32). One case relapsed at after-treatment week 32. Next-generation sequencing (NGS) was conducted and showed that the treatment failure case had two suspected antiviral resistance mutations, NS5A V28M (a change of V to M at position 28) and NS5B A442V, compared with the baseline. Overall, this newly identified 6xj subtype further confirmed the high diversity of the HCV-6 genotype. The newly identified resistance-associated amino acid substitutions may help inform future clinical treatments. IMPORTANCE This study investigated the genetic diversity of hepatitis C virus (HCV), particularly in relation to genotype 6, which is prevalent in Yunnan, China, and is often difficult to treat successfully. We identified a new HCV-6 subtype, 6xj, which is an ancient strain. Moreover, all eight individuals with the novel subtype received follow-up assessments at 44 weeks from the beginning of their treatments. One case relapsed after 8 months of withdrawal. NGS was conducted and showed that the isolate from the treatment failure case had two suspected antiviral resistance mutations, NS5A V28M and NS5B A442V, compared with the baseline. Overall, this newly identified 6xj subtype further confirmed the high diversity of the HCV-6 genotype. The newly identified resistance-associated amino acid substitutions may help inform future clinical treatments. We believe that our study makes a significant contribution to the literature based on the results described above.
Keywords: DAA; RAS; follow-up; hepatitis C virus; novel subtype; resistance-associated substitutions.
Figures

Similar articles
-
HCV Genotype 6a Escape From and Resistance to Velpatasvir, Pibrentasvir, and Sofosbuvir in Robust Infectious Cell Culture Models.Gastroenterology. 2018 Jun;154(8):2194-2208.e12. doi: 10.1053/j.gastro.2018.02.017. Epub 2018 Feb 15. Gastroenterology. 2018. PMID: 29454794
-
Resistance analysis in patients with genotype 1-6 HCV infection treated with sofosbuvir/velpatasvir in the phase III studies.J Hepatol. 2018 May;68(5):895-903. doi: 10.1016/j.jhep.2017.11.032. Epub 2017 Dec 6. J Hepatol. 2018. PMID: 29221887 Clinical Trial.
-
Hepatitis C virus diversity and treatment outcomes in Benin: a prospective cohort study.Lancet Microbe. 2024 Jul;5(7):697-706. doi: 10.1016/S2666-5247(24)00041-7. Epub 2024 Jun 15. Lancet Microbe. 2024. PMID: 38889738
-
Sofosbuvir-velpatasvir: A single-tablet treatment for hepatitis C infection of all genotypes.Am J Health Syst Pharm. 2017 Jul 15;74(14):1045-1052. doi: 10.2146/ajhp60632. Am J Health Syst Pharm. 2017. PMID: 28687550 Review.
-
Sofosbuvir/Velpatasvir/Voxilaprevir: A Review in Chronic Hepatitis C.Drugs. 2018 Apr;78(5):577-587. doi: 10.1007/s40265-018-0895-5. Drugs. 2018. PMID: 29546556 Review.
Cited by
-
The distribution of hepatitis C viral genotypes shifted among chronic hepatitis C patients in Yunnan, China, between 2008-2018.Front Cell Infect Microbiol. 2023 Jul 11;13:1092936. doi: 10.3389/fcimb.2023.1092936. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 37496804 Free PMC article.
-
Treatment Response and Drug Resistance Profiling of Genotype 6 of Hepatitis C Virus in HCV/HIV Co-Infected Patients: A Pilot Study from INDIA.Viruses. 2022 Apr 30;14(5):944. doi: 10.3390/v14050944. Viruses. 2022. PMID: 35632686 Free PMC article.
-
Genotype Distribution and Migration Patterns of Hepatitis C Virus in Shandong Province, China: Molecular Epidemiology and Phylogenetic Study.JMIR Med Inform. 2025 Aug 18;13:e60207. doi: 10.2196/60207. JMIR Med Inform. 2025. PMID: 40825211 Free PMC article.
References
-
- Soria ME, García-Crespo C, Martínez-González B, Vázquez-Sirvent L, Lobo-Vega R, de Ávila AI, Gallego I, Chen Q, García-Cehic D, Llorens-Revull M, Briones C, Gómez J, Ferrer-Orta C, Verdaguer N, Gregori J, Rodríguez-Frías F, Buti M, Esteban JI, Domingo E, Quer J, Perales C. 2020. Amino acid substitutions associated with treatment failure of hepatitis C virus infection. J Clin Microbiol 58:e01985-20. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

