Evolution of Methyltransferase-Like (METTL) Proteins in Metazoa: A Complex Gene Family Involved in Epitranscriptomic Regulation and Other Epigenetic Processes
- PMID: 34480573
- PMCID: PMC8662637
- DOI: 10.1093/molbev/msab267
Evolution of Methyltransferase-Like (METTL) Proteins in Metazoa: A Complex Gene Family Involved in Epitranscriptomic Regulation and Other Epigenetic Processes
Abstract
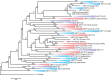
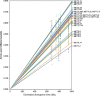
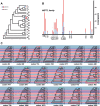
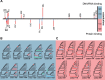
The methyltransferase-like (METTL) proteins constitute a family of seven-beta-strand methyltransferases with S-adenosyl methionine-binding domains that modify DNA, RNA, and proteins. Methylation by METTL proteins contributes to the epigenetic, and in the case of RNA modifications, epitranscriptomic regulation of a variety of biological processes. Despite their functional importance, most investigations of the substrates and functions of METTLs within metazoans have been restricted to model vertebrate taxa. In the present work, we explore the evolutionary mechanisms driving the diversification and functional differentiation of 33 individual METTL proteins across Metazoa. Our results show that METTLs are nearly ubiquitous across the animal kingdom, with most having arisen early in metazoan evolution (i.e., occur in basal metazoan phyla). Individual METTL lineages each originated from single independent ancestors, constituting monophyletic clades, which suggests that each METTL was subject to strong selective constraints driving its structural and/or functional specialization. Interestingly, a similar process did not extend to the differentiation of nucleoside-modifying and protein-modifying METTLs (i.e., each METTL type did not form a unique monophyletic clade). The members of these two types of METTLs also exhibited differences in their rates of evolution. Overall, we provide evidence that the long-term evolution of METTL family members was driven by strong purifying selection, which in combination with adaptive selection episodes, led to the functional specialization of individual METTL lineages. This work contributes useful information regarding the evolution of a gene family that fulfills a variety of epigenetic functions, and can have profound influences on molecular processes and phenotypic traits.
Keywords: METTL; epigenetics; metazoan; methyltransferase; phylogenetics; selection.
© The Author(s) 2021. Published by Oxford University Press on behalf of the Society for Molecular Biology and Evolution.
Figures


References
-
- Angulo JF, Mauffirey P, Pinon-Lataillade G, Miccoli L, Biard DSF.. 2005. Putative roles of kin17, a mammalian protein binding curved DNA, in transcription. In: Ohyama T, editor. DNA conformation and transcription. Molecular Biology Intelligence Unit. Boston (MA): Springer. p. 75–89.
-
- Bhattacharyya M, De S, Chakrabarti S.. 2020. Origin and evolution of DNA methyltransferases (DNMT) along the tree of life: a multi-genome survey. bioRxiv 2020.04.09.033167.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

