Microarray Analysis Identifies Key Differentially Expressed Circular RNAs in Aged Mice With Postoperative Cognitive Dysfunction
- PMID: 34483886
- PMCID: PMC8415796
- DOI: 10.3389/fnagi.2021.716383
Microarray Analysis Identifies Key Differentially Expressed Circular RNAs in Aged Mice With Postoperative Cognitive Dysfunction
Abstract
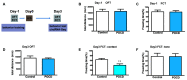
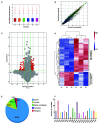
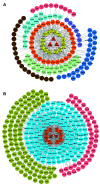
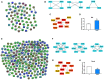
Postoperative cognitive dysfunction (POCD) is a common complication in elderly patients. Circular RNAs (circRNAs) may contribute to neurodegenerative diseases. However, the role of circRNAs in POCD in aged mice has not yet been reported. This study aimed to explore the potential circRNAs in a POCD model. First, a circRNA microarray was used to analyze the expression profiles. Differentially expressed circRNAs were validated using quantitative real-time polymerase chain reaction. A bioinformatics analysis was then used to construct a competing endogenous RNA (ceRNA) network. The database for annotation, visualization, and integrated discovery was used to perform Gene Ontology (GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment analysis of circRNA-related genes. Moreover, protein-protein interactions were analyzed to predict the circRNA-regulated hub genes using the STRING and molecular complex detection plug-in of Cytoscape. Microarray screen 124 predicted circRNAs in the POCD of aged mice. We found that the up/downregulated circRNAs were involved in multiple signaling pathways. Hub genes, including Egfr and Prkacb, were identified and may be regulated by ceRNA networks. These results suggest that circRNAs are dysexpressed in the hippocampus and may contribute to POCD in aged mice.
Keywords: ceRNA network; circRNAs; expression profile; miRNAs; postoperative cognitive dysfunction.
Copyright © 2021 Wu, Liu, Wang, Chen, Huang, Sun, Ma, Wan, Sun and Miao.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Identification of the Potential Key Circular RNAs in Elderly Patients With Postoperative Cognitive Dysfunction.Front Aging Neurosci. 2020 Jun 23;12:165. doi: 10.3389/fnagi.2020.00165. eCollection 2020. Front Aging Neurosci. 2020. PMID: 32655392 Free PMC article.
-
Characterization of circRNA-Associated-ceRNA Networks Involved in the Pathogenesis of Postoperative Cognitive Dysfunction in Aging Mice.Front Aging Neurosci. 2022 Apr 4;14:727805. doi: 10.3389/fnagi.2022.727805. eCollection 2022. Front Aging Neurosci. 2022. PMID: 35444525 Free PMC article.
-
Abnormal expression of circRNA_089763 in the plasma exosomes of patients with post‑operative cognitive dysfunction after coronary artery bypass grafting.Mol Med Rep. 2019 Sep;20(3):2549-2562. doi: 10.3892/mmr.2019.10521. Epub 2019 Jul 24. Mol Med Rep. 2019. PMID: 31524256 Free PMC article.
-
Construction of Circular RNA-MicroRNA-Messenger RNA Regulatory Network of Recurrent Implantation Failure to Explore Its Potential Pathogenesis.Front Genet. 2021 Feb 16;11:627459. doi: 10.3389/fgene.2020.627459. eCollection 2020. Front Genet. 2021. PMID: 33664765 Free PMC article.
-
Recent progress on the role of non-coding RNA in postoperative cognitive dysfunction.Front Cell Neurosci. 2022 Oct 13;16:1024475. doi: 10.3389/fncel.2022.1024475. eCollection 2022. Front Cell Neurosci. 2022. PMID: 36313620 Free PMC article. Review.
Cited by
-
Research progress on perioperative blood-brain barrier damage and its potential mechanism.Front Cell Dev Biol. 2023 Apr 10;11:1174043. doi: 10.3389/fcell.2023.1174043. eCollection 2023. Front Cell Dev Biol. 2023. PMID: 37101615 Free PMC article. Review.
-
Decoding competitive endogenous RNA regulatory network in postoperative cognitive dysfunction.Front Neurosci. 2022 Sep 20;16:972918. doi: 10.3389/fnins.2022.972918. eCollection 2022. Front Neurosci. 2022. PMID: 36203795 Free PMC article.
-
Identification of Potential Key circRNAs in Aged Mice With Postoperative Delirium.Front Mol Neurosci. 2022 Apr 14;15:836534. doi: 10.3389/fnmol.2022.836534. eCollection 2022. Front Mol Neurosci. 2022. PMID: 35493320 Free PMC article.
-
Novel implications of a strictly monomorphic (GCC) repeat in the human PRKACB gene.Sci Rep. 2021 Oct 19;11(1):20629. doi: 10.1038/s41598-021-99932-3. Sci Rep. 2021. PMID: 34667254 Free PMC article.
-
Identification of circRNA-miRNA-mRNA networks to explore the molecular mechanism and immune regulation of postoperative neurocognitive disorder.Aging (Albany NY). 2022 Oct 21;14(20):8374-8393. doi: 10.18632/aging.204348. Epub 2022 Oct 21. Aging (Albany NY). 2022. PMID: 36279395 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

