PIWI silencing mechanism involving the retrotransposon nimbus orchestrates resistance to infection with Schistosoma mansoni in the snail vector, Biomphalaria glabrata
- PMID: 34495959
- PMCID: PMC8462715
- DOI: 10.1371/journal.pntd.0009094
PIWI silencing mechanism involving the retrotransposon nimbus orchestrates resistance to infection with Schistosoma mansoni in the snail vector, Biomphalaria glabrata
Abstract
Background: Schistosomiasis remains widespread in many regions despite efforts at its elimination. By examining changes in the transcriptome at the host-pathogen interface in the snail Biomphalaria glabrata and the blood fluke Schistosoma mansoni, we previously demonstrated that an early stress response in juvenile snails, manifested by induction of heat shock protein 70 (Hsp 70) and Hsp 90 and of the reverse transcriptase (RT) domain of the B. glabrata non-LTR- retrotransposon, nimbus, were critical for B. glabrata susceptibility to S. mansoni. Subsequently, juvenile B. glabrata BS-90 snails, resistant to S. mansoni at 25°C become susceptible by the F2 generation when maintained at 32°C, indicating an epigenetic response.
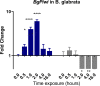
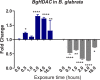
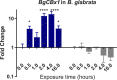
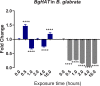
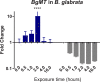
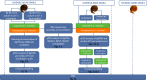
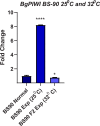
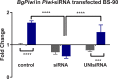
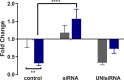
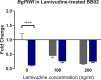
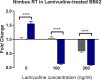
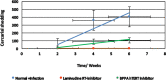
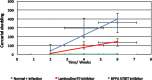
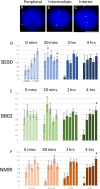
Methodology/principal findings: To better understand this plasticity in susceptibility of the BS-90 snail, mRNA sequences were examined from S. mansoni exposed juvenile BS-90 snails cultured either at 25°C (non-permissive temperature) or 32°C (permissive). Comparative analysis of transcriptomes from snails cultured at the non-permissive and permissive temperatures revealed that whereas stress related transcripts dominated the transcriptome of susceptible BS-90 juvenile snails at 32°C, transcripts encoding proteins with a role in epigenetics, such as PIWI (BgPiwi), chromobox protein homolog 1 (BgCBx1), histone acetyltransferase (BgHAT), histone deacetylase (BgHDAC) and metallotransferase (BgMT) were highly expressed in those cultured at 25°C. To identify robust candidate transcripts that will underscore the anti-schistosome phenotype in B. glabrata, further validation of the differential expression of the above transcripts was performed by using the resistant BS-90 (25°C) and the BBO2 susceptible snail stock whose genome has now been sequenced and represents an invaluable resource for molecular studies in B. glabrata. A role for BgPiwi in B. glabrata susceptibility to S. mansoni, was further examined by using siRNA corresponding to the BgPiwi encoding transcript to suppress expression of BgPiwi, rendering the resistant BS-90 juvenile snail susceptible to infection at 25°C. Given transposon silencing activity of PIWI as a facet of its role as guardian of the integrity of the genome, we examined the expression of the nimbus RT encoding transcript at 120 min after infection of resistant BS90 piwi-siRNA treated snails. We observed that nimbus RT was upregulated, indicating that modulation of the transcription of the nimbus RT was associated with susceptibility to S. mansoni in BgPiwi-siRNA treated BS-90 snails. Furthermore, treatment of susceptible BBO2 snails with the RT inhibitor lamivudine, before exposure to S. mansoni, blocked S. mansoni infection concurrent with downregulation of the nimbus RT transcript and upregulation of the BgPiwi encoding transcript in the lamivudine-treated, schistosome-exposed susceptible snails.
Conclusions and significance: These findings support a role for the interplay of BgPiwi and nimbus in the epigenetic modulation of plasticity of resistance/susceptibility in the snail-schistosome relationship.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures
Similar articles
-
Reversing the resistance phenotype of the Biomphalaria glabrata snail host Schistosoma mansoni infection by temperature modulation.PLoS Pathog. 2012;8(4):e1002677. doi: 10.1371/journal.ppat.1002677. Epub 2012 Apr 26. PLoS Pathog. 2012. PMID: 22577362 Free PMC article.
-
Identification of immediate response genes dominantly expressed in juvenile resistant and susceptible Biomphalaria glabrata snails upon exposure to Schistosoma mansoni.Mol Biochem Parasitol. 2010 Jan;169(1):27-39. doi: 10.1016/j.molbiopara.2009.09.009. Epub 2009 Oct 6. Mol Biochem Parasitol. 2010. PMID: 19815034 Free PMC article.
-
Susceptibility of Snails to Infection with Schistosomes is influenced by Temperature and Expression of Heat Shock Proteins.Epidemiology (Sunnyvale). 2015 Jun;5(2):189. doi: 10.4172/2161-1165.1000189. Epub 2015 Jun 21. Epidemiology (Sunnyvale). 2015. PMID: 26504668 Free PMC article.
-
Epigenetic modulation, stress and plasticity in susceptibility of the snail host, Biomphalaria glabrata, to Schistosoma mansoni infection.Int J Parasitol. 2016 Jun;46(7):389-94. doi: 10.1016/j.ijpara.2016.03.003. Epub 2016 Apr 4. Int J Parasitol. 2016. PMID: 27056272 Review.
-
Are Biomphalaria snails resistant to Schistosoma mansoni?J Helminthol. 2005 Sep;79(3):187-91. doi: 10.1079/joh2005299. J Helminthol. 2005. PMID: 16153311 Review.
Cited by
-
Genotypic-specific heat shock response of vector susceptibility to Schistosoma mansoni.Ecosphere. 2022 Aug;13(8):e4207. doi: 10.1002/ecs2.4207. Epub 2022 Aug 18. Ecosphere. 2022. PMID: 36590709 Free PMC article.
-
Introduction of epigenetic variation contribute to resistance against the human parasite Schistosoma mansoni.Epigenetics Chromatin. 2025 Jul 14;18(1):43. doi: 10.1186/s13072-025-00607-4. Epigenetics Chromatin. 2025. PMID: 40660336 Free PMC article.
-
Exploring the immune interactions between Oncomelania hupensis and Schistosoma japonicum, with a cross-comparison of immunological research progress in other intermediate host snails.Parasit Vectors. 2023 Dec 13;16(1):453. doi: 10.1186/s13071-023-06011-9. Parasit Vectors. 2023. PMID: 38093363 Free PMC article. Review.
-
An Overview of Transcriptional Responses of Schistosome-Susceptible (M line) or -Resistant (BS-90) Biomphalaria glabrata Exposed or Not to Schistosoma mansoni Infection.Front Immunol. 2022 Jan 12;12:805882. doi: 10.3389/fimmu.2021.805882. eCollection 2021. Front Immunol. 2022. PMID: 35095891 Free PMC article.
References
-
- Knopp S, Ame SM, Hattendorf J, Ali SM, Khamis IS, Bakar F, et al.. Urogenital schistosomiasis elimination in Zanzibar: accuracy of urine filtration and haematuria reagent strips for diagnosing light intensity Schistosoma haematobium infections. Parasit Vectors. 2018;11(1):552. Epub 2018/10/26. doi: 10.1186/s13071-018-3136-6; PubMed Central PMCID: PMC6199745. - DOI - PMC - PubMed
-
- Shen Y, Wiegand RE, Olsen A, King CH, Kittur N, Binder S, et al.. Five-Year Impact of Different Multi-Year Mass Drug Administration Strategies on Childhood Schistosoma mansoni-Associated Morbidity: A Combined Analysis from the Schistosomiasis Consortium for Operational Research and Evaluation Cohort Studies in the Lake Victoria Regions of Kenya and Tanzania. Am J Trop Med Hyg. 2019;101(6):1336–44. Epub 2019/08/14. doi: 10.4269/ajtmh.19-0273 ; PubMed Central PMCID: PMC6896894. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

