The Molecular Mechanism of Multiple Organ Dysfunction and Targeted Intervention of COVID-19 Based on Time-Order Transcriptomic Analysis
- PMID: 34504502
- PMCID: PMC8421734
- DOI: 10.3389/fimmu.2021.729776
The Molecular Mechanism of Multiple Organ Dysfunction and Targeted Intervention of COVID-19 Based on Time-Order Transcriptomic Analysis
Abstract
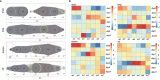
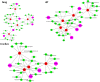
Coronavirus disease 2019 (COVID-19) pandemic is caused by the novel coronavirus that has spread rapidly around the world, leading to high mortality because of multiple organ dysfunction; however, its underlying molecular mechanism is unknown. To determine the molecular mechanism of multiple organ dysfunction, a bioinformatics analysis method based on a time-order gene co-expression network (TO-GCN) was performed. First, gene expression profiles were downloaded from the gene expression omnibus database (GSE161200), and a TO-GCN was constructed using the breadth-first search (BFS) algorithm to infer the pattern of changes in the different organs over time. Second, Gene Ontology enrichment analysis was used to analyze the main biological processes related to COVID-19. The initial gene modules for the immune response of different organs were defined as the research object. The STRING database was used to construct a protein-protein interaction network of immune genes in different organs. The PageRank algorithm was used to identify five hub genes in each organ. Finally, the Comparative Toxicogenomics Database played an important role in exploring the potential compounds that target the hub genes. The results showed that there were two types of biological processes: the body's stress response and cell-mediated immune response involving the lung, trachea, and olfactory bulb (olf) after being infected by COVID-19. However, a unique biological process related to the stress response is the regulation of neuronal signals in the brain. The stress response was heterogeneous among different organs. In the lung, the regulation of DNA morphology, angiogenesis, and mitochondrial-related energy metabolism are specific biological processes related to the stress response. In particular, an effect on tracheal stress response was made by the regulation of protein metabolism and rRNA metabolism-related biological processes, as biological processes. In the olf, the distinctive stress responses consist of neural signal transmission and brain behavior. In addition, myeloid leukocyte activation and myeloid leukocyte-mediated immunity in response to COVID-19 can lead to a cytokine storm. Immune genes such as SRC, RHOA, CD40LG, CSF1, TNFRSF1A, FCER1G, ICAM1, LAT, LCN2, PLAU, CXCL10, ICAM1, CD40, IRF7, and B2M were predicted to be the hub genes in the cytokine storm. Furthermore, we inferred that resveratrol, acetaminophen, dexamethasone, estradiol, statins, curcumin, and other compounds are potential target drugs in the treatment of COVID-19.
Keywords: BFS algorithm; COVID-19; TO-GCN; cytokine storm; multiple organ dysfunction; multiple organ heterogeneity; targeted intervention.
Copyright © 2021 Zou, Su, Wang, Yi, Qiu, Yin, Zhou, Niu, Wang and Su.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures





References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

