The gut microbiome in konzo
- PMID: 34508085
- PMCID: PMC8433213
- DOI: 10.1038/s41467-021-25694-1
The gut microbiome in konzo
Abstract
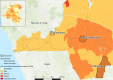
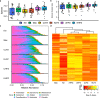
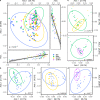
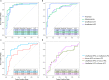
Konzo, a distinct upper motor neuron disease associated with a cyanogenic diet and chronic malnutrition, predominately affects children and women of childbearing age in sub-Saharan Africa. While the exact biological mechanisms that cause this disease have largely remained elusive, host-genetics and environmental components such as the gut microbiome have been implicated. Using a large study population of 180 individuals from the Democratic Republic of the Congo, where konzo is most frequent, we investigate how the structure of the gut microbiome varied across geographical contexts, as well as provide the first insight into the gut flora of children affected with this debilitating disease using shotgun metagenomic sequencing. Our findings indicate that the gut microbiome structure is highly variable depending on region of sampling, but most interestingly, we identify unique enrichments of bacterial species and functional pathways that potentially modulate the susceptibility of konzo in prone regions of the Congo.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures
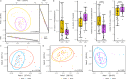

Comment in
-
Gut microbiome composition linked to konzo risk.Nat Rev Neurol. 2021 Nov;17(11):660. doi: 10.1038/s41582-021-00572-y. Nat Rev Neurol. 2021. PMID: 34594002 No abstract available.

