Correction to: Post-diapause transcriptomic restarts: insight from a high-latitude copepod
- PMID: 34521338
- PMCID: PMC8439028
- DOI: 10.1186/s12864-021-07868-9
Correction to: Post-diapause transcriptomic restarts: insight from a high-latitude copepod
Figures



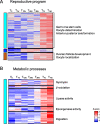
Erratum for
-
Post-diapause transcriptomic restarts: insight from a high-latitude copepod.BMC Genomics. 2021 Jun 3;22(1):409. doi: 10.1186/s12864-021-07557-7. BMC Genomics. 2021. PMID: 34082716 Free PMC article.
References
-
- Desgraupes B. clusterCrit: Clustering Indices. R package version 1.2. 6. 2015.
Publication types
LinkOut - more resources
Full Text Sources

