Characterization of variants of uncertain significance in isovaleryl-CoA dehydrogenase identified through newborn screening: An approach for faster analysis
- PMID: 34535384
- PMCID: PMC8578405
- DOI: 10.1016/j.ymgme.2021.08.012
Characterization of variants of uncertain significance in isovaleryl-CoA dehydrogenase identified through newborn screening: An approach for faster analysis
Abstract
Introduction: Clinical standard of care for newborn screening (NBS) is acylcarnitine metabolites quantitation by tandem mass spectrometry (MS/MS) from dried blood spots. Follow up sequencing often results in identification of one or more variants of uncertain significance (VUS). Isovaleric acidemia (IVA) is an autosomal recessive inborn error of metabolism caused by deficiency of isovaleryl-CoA dehydrogenase (IVDH) in the Leu catabolism pathway. Many IVD mutations are characterized as VUS complicating IVA clinical diagnoses and treatment. We present a testing platform approach to confirm the functional implication of VUS identified in newborns with IVA applicable to multiple inborn errors of metabolism identified by NBS.
Methods: An IVD null HEK293T cell culture model was generated by using a dual sgRNA CRISPR/Cas9 genome-editing strategy targeting IVD exons 2-3. Clonal cell lines were confirmed by a combination of genomic breakpoint sequencing and droplet digital PCR. The IVD null model had no IVDH antigen signal and 96% reduction in IVDH enzyme activity. The IVD null model was transfected with vectors containing control or variant IVD and functional assays were performed to determine variant pathogenicity.
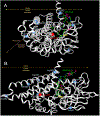
Results: c.149G > C (p.Arg50Pro; precursor numbering), c.986T > C (p.Met329Thr), and c.1010G > A (p.Arg337Gln), c.1179del394 f. mutant proteins had reduced IVDH protein and activity. c.932C > T (p.Ala311Val), c.707C > T (p.Thr236Ile), and c.1232G > A (p.Arg411Gln) had stable IVDH protein, but no enzyme activity. c.521T > G (p.Val174Gly) had normal IVDH protein and activity. IVD variant transfection results confirmed results from IVA fibroblasts containing the same variants.
Conclusions: We have developed an IVD null HEK293T cell line to rapidly allow determination of VUS pathogenicity following identification of novel alleles by clinical sequencing following positive NBS results for suspected IVA. We suggest similar models can be generated via genome-editing for high throughput assessment of VUS function for a multitude of inborn errors of metabolism and can ideally supplement NBS programs.
Keywords: Isovaleric acidemia; Isovaleryl-CoA dehydrogenase; Newborn screening; Organic acidemia; Variants of uncertain significance.
Copyright © 2021 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest The authors certify that they have no affiliations with or involvement in any organization or entity with any financial or non-financial interest in the subject matter or materials discussed in this manuscript.
Figures

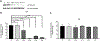
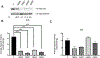

Similar articles
-
Prevalence and mutation analysis of short/branched chain acyl-CoA dehydrogenase deficiency (SBCADD) detected on newborn screening in Wisconsin.Mol Genet Metab. 2013 Sep-Oct;110(1-2):111-5. doi: 10.1016/j.ymgme.2013.03.021. Epub 2013 Apr 15. Mol Genet Metab. 2013. PMID: 23712021 Free PMC article.
-
Phenotypic Variability and Newly Identified Mutations of the IVD Gene in Japanese Patients with Isovaleric Acidemia.Tohoku J Exp Med. 2015 Jun;236(2):103-6. doi: 10.1620/tjem.236.103. Tohoku J Exp Med. 2015. PMID: 26018748
-
Newborn screening for isovaleric acidemia in Quanzhou, China.Clin Chim Acta. 2020 Oct;509:25-29. doi: 10.1016/j.cca.2020.06.010. Epub 2020 Jun 4. Clin Chim Acta. 2020. PMID: 32505769
-
Isovaleric acidemia: new aspects of genetic and phenotypic heterogeneity.Am J Med Genet C Semin Med Genet. 2006 May 15;142C(2):95-103. doi: 10.1002/ajmg.c.30089. Am J Med Genet C Semin Med Genet. 2006. PMID: 16602101 Free PMC article. Review.
-
Aspects of Newborn Screening in Isovaleric Acidemia.Int J Neonatal Screen. 2018 Jan 29;4(1):7. doi: 10.3390/ijns4010007. eCollection 2018 Mar. Int J Neonatal Screen. 2018. PMID: 33072933 Free PMC article. Review.
Cited by
-
Functional analysis of a novel homozygous missense IVD gene variant: a case report with dual genetic diagnoses.Front Pediatr. 2025 Feb 10;13:1494530. doi: 10.3389/fped.2025.1494530. eCollection 2025. Front Pediatr. 2025. PMID: 39995896 Free PMC article.
-
Structural Insights into Isovaleryl-Coenzyme A Dehydrogenase: Mechanisms of Substrate Specificity and Implications of Isovaleric Acidemia-Associated Mutations.Research (Wash D C). 2025 May 28;8:0661. doi: 10.34133/research.0661. eCollection 2025. Research (Wash D C). 2025. PMID: 40851894 Free PMC article.
-
Hypoglycaemia Metabolic Gene Panel Testing.Front Endocrinol (Lausanne). 2022 Mar 29;13:826167. doi: 10.3389/fendo.2022.826167. eCollection 2022. Front Endocrinol (Lausanne). 2022. PMID: 35422763 Free PMC article. Review.
-
Characterization of exonic variants of uncertain significance in very long-chain acyl-CoA dehydrogenase identified through newborn screening.J Inherit Metab Dis. 2022 May;45(3):529-540. doi: 10.1002/jimd.12492. Epub 2022 Mar 11. J Inherit Metab Dis. 2022. PMID: 35218577 Free PMC article.
References
-
- Oglesbee D, et al., Second-tier test for quantification of alloisoleucine and branched-chain amino acids in dried blood spots to improve newborn screening for maple syrup urine disease (MSUD). Clin Chem, 2008. 54(3): p. 542–9. - PubMed
-
- Narravula A, et al., Variants of uncertain significance in newborn screening disorders: implications for large-scale genomic sequencing. Genet Med, 2017. 19(1): p. 77–82. - PubMed
-
- Wilcken B, Medicine. Newborn screening: gaps in the evidence. Science, 2013. 342(6155): p. 197–8. - PubMed
-
- Online, M.I.o.M., OMIM Isovaleric Acidemia, IVA #243500. OMIM, Mendelian Inheritance of Man, OMIM.
Publication types
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

