A 584 bp deletion in CTRB2 inhibits chymotrypsin B2 activity and secretion and confers risk of pancreatic cancer
- PMID: 34559995
- PMCID: PMC8546220
- DOI: 10.1016/j.ajhg.2021.09.002
A 584 bp deletion in CTRB2 inhibits chymotrypsin B2 activity and secretion and confers risk of pancreatic cancer
Abstract
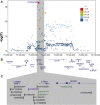
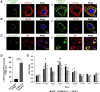
Genome-wide association studies (GWASs) have discovered 20 risk loci in the human genome where germline variants associate with risk of pancreatic ductal adenocarcinoma (PDAC) in populations of European ancestry. Here, we fine-mapped one such locus on chr16q23.1 (rs72802365, p = 2.51 × 10-17, OR = 1.36, 95% CI = 1.31-1.40) and identified colocalization (PP = 0.87) with aberrant exon 5-7 CTRB2 splicing in pancreatic tissues (pGTEx = 1.40 × 10-69, βGTEx = 1.99; pLTG = 1.02 × 10-30, βLTG = 1.99). Imputation of a 584 bp structural variant overlapping exon 6 of CTRB2 into the GWAS datasets resulted in a highly significant association with pancreatic cancer risk (p = 2.83 × 10-16, OR = 1.36, 95% CI = 1.31-1.42), indicating that it may underlie this signal. Exon skipping attributable to the deletion (risk) allele introduces a premature stop codon in exon 7 of CTRB2, yielding a truncated chymotrypsinogen B2 protein that lacks chymotrypsin activity, is poorly secreted, and accumulates intracellularly in the endoplasmic reticulum (ER). We propose that intracellular accumulation of a nonfunctional chymotrypsinogen B2 protein leads to ER stress and pancreatic inflammation, which may explain the increased pancreatic cancer risk in carriers of CTRB2 exon 6 deletion alleles.
Keywords: genetics; pancreatic cancer; pancreatic enzymes; polymorphic variation; splicing QTL.
Published by Elsevier Inc.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures


References
-
- Siegel R.L., Miller K.D., Jemal A. Cancer statistics, 2019. CA Cancer J. Clin. 2019;69:7–34. - PubMed
-
- Ferlay J., Colombet M., Soerjomataram I., Dyba T., Randi G., Bettio M., Gavin A., Visser O., Bray F. Cancer incidence and mortality patterns in Europe: Estimates for 40 countries and 25 major cancers in 2018. Eur. J. Cancer. 2018;103:356–387. - PubMed
-
- Rahib L., Smith B.D., Aizenberg R., Rosenzweig A.B., Fleshman J.M., Matrisian L.M. Projecting cancer incidence and deaths to 2030: the unexpected burden of thyroid, liver, and pancreas cancers in the United States. Cancer Res. 2014;74:2913–2921. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Miscellaneous

