Intertwined and Finely Balanced: Endoplasmic Reticulum Morphology, Dynamics, Function, and Diseases
- PMID: 34571990
- PMCID: PMC8472773
- DOI: 10.3390/cells10092341
Intertwined and Finely Balanced: Endoplasmic Reticulum Morphology, Dynamics, Function, and Diseases
Abstract
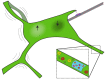
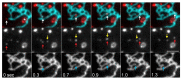
The endoplasmic reticulum (ER) is an organelle that is responsible for many essential subcellular processes. Interconnected narrow tubules at the periphery and thicker sheet-like regions in the perinuclear region are linked to the nuclear envelope. It is becoming apparent that the complex morphology and dynamics of the ER are linked to its function. Mutations in the proteins involved in regulating ER structure and movement are implicated in many diseases including neurodegenerative diseases such as Alzheimer's, Parkinson's, and amyotrophic lateral sclerosis (ALS). The ER is also hijacked by pathogens to promote their replication. Bacteria such as Legionella pneumophila and Chlamydia trachomatis, as well as the Zika virus, bind to ER morphology and dynamics-regulating proteins to exploit the functions of the ER to their advantage. This review covers our understanding of ER morphology, including the functional subdomains and membrane contact sites that the organelle forms. We also focus on ER dynamics and the current efforts to quantify ER motion and discuss the diseases related to ER morphology and dynamics.
Keywords: anomalous diffusion; dynamics; dynein; endoplasmic reticulum (ER); kinesin; membrane contact site (MCS); microtubule; morphology.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the writing of the manuscript, or in the decision to publish the results.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

