Quantitative Approaches to Study Retinal Neurogenesis
- PMID: 34572408
- PMCID: PMC8471905
- DOI: 10.3390/biomedicines9091222
Quantitative Approaches to Study Retinal Neurogenesis
Abstract
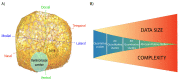
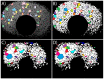
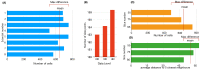
The study of the development of the vertebrate retina can be addressed from several perspectives: from a purely qualitative to a more quantitative approach that takes into account its spatio-temporal features, its three-dimensional structure and also the regulation and properties at the systems level. Here, we review the ongoing transition toward a full four-dimensional characterization of the developing vertebrate retina, focusing on the challenges at the experimental, image acquisition, image processing and quantification. Using the developing zebrafish retina, we illustrate how quantitative data extracted from these type of highly dense, three-dimensional tissues depend strongly on the image quality, image processing and algorithms used to segment and quantify. Therefore, we propose that the scientific community that focuses on developmental systems could strongly benefit from a more detailed disclosure of the tools and pipelines used to process and analyze images from biological samples.
Keywords: imaging; quantitative biology; retinogenesis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
A dynamic and expandable digital 3D-atlas maker for monitoring the temporal changes in tissue growth during hindbrain morphogenesis.Elife. 2022 Sep 28;11:e78300. doi: 10.7554/eLife.78300. Elife. 2022. PMID: 36169400 Free PMC article.
-
Live imaging of developing mouse retinal slices.Neural Dev. 2018 Sep 15;13(1):23. doi: 10.1186/s13064-018-0120-y. Neural Dev. 2018. PMID: 30219109 Free PMC article.
-
Unifying developmental programs for embryonic and postembryonic neurogenesis in the zebrafish retina.Development. 2020 Jun 19;147(12):dev185660. doi: 10.1242/dev.185660. Development. 2020. PMID: 32467236
-
Spatial and spatio-temporal statistical analyses of retinal images: a review of methods and applications.BMJ Open Ophthalmol. 2020 May 28;5(1):e000479. doi: 10.1136/bmjophth-2020-000479. eCollection 2020. BMJ Open Ophthalmol. 2020. PMID: 32537517 Free PMC article. Review.
-
Expression and function of the LIM-homeodomain transcription factor Islet-1 in the developing and mature vertebrate retina.Exp Eye Res. 2015 Sep;138:22-31. doi: 10.1016/j.exer.2015.06.021. Epub 2015 Jun 26. Exp Eye Res. 2015. PMID: 26122047 Review.
References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

