The Tumor Microenvironment-Dependent Transcription Factors AHR and HIF-1α Are Dispensable for Leukemogenesis in the Eµ-TCL1 Mouse Model of Chronic Lymphocytic Leukemia
- PMID: 34572746
- PMCID: PMC8466120
- DOI: 10.3390/cancers13184518
The Tumor Microenvironment-Dependent Transcription Factors AHR and HIF-1α Are Dispensable for Leukemogenesis in the Eµ-TCL1 Mouse Model of Chronic Lymphocytic Leukemia
Abstract
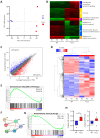
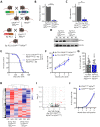
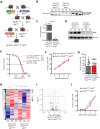
Chronic lymphocytic leukemia (CLL) is the most frequent leukemia in the elderly and is characterized by the accumulation of mature B lymphocytes in peripheral blood and primary lymphoid organs. In order to proliferate, leukemic cells are highly dependent on complex interactions with their microenvironment in proliferative niches. Not only soluble factors and BCR stimulation are important for their survival and proliferation, but also the activation of transcription factors through different signaling pathways. The aryl hydrocarbon receptor (AHR) and hypoxia-inducible factor (HIF)-1α are two transcription factors crucial for cancer development, whose activities are dependent on tumor microenvironment conditions, such as the presence of metabolites from the tryptophan pathway and hypoxia, respectively. In this study, we addressed the potential role of AHR and HIF-1α in chronic lymphocytic leukemia (CLL) development in vivo. To this end, we crossed the CLL mouse model Eµ-TCL1 with the corresponding transcription factor-conditional knock-out mice to delete one or both transcription factors in CD19+ B cells only. Despite AHR and HIF-1α being activated in CLL cells, deletion of either or both of them had no impact on CLL progression or survival in vivo, suggesting that these transcription factors are not crucial for leukemogenesis in CLL.
Keywords: AHR; HIF1α; chronic lymphocytic leukemia; tumor microenvironment.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures

Similar articles
-
Lessons learned from the Eµ-TCL1 mouse model of CLL.Semin Hematol. 2024 Jun;61(3):194-200. doi: 10.1053/j.seminhematol.2024.05.002. Epub 2024 May 10. Semin Hematol. 2024. PMID: 38839457 Review.
-
HIF-1α regulates the interaction of chronic lymphocytic leukemia cells with the tumor microenvironment.Blood. 2016 Apr 21;127(16):1987-97. doi: 10.1182/blood-2015-07-657056. Epub 2016 Jan 29. Blood. 2016. PMID: 26825709 Free PMC article.
-
CD74 is dispensable for development of chronic lymphocytic leukemia in Eµ-TCL1 transgenic mice.Leuk Lymphoma. 2020 Dec;61(12):2799-2810. doi: 10.1080/10428194.2020.1791851. Epub 2020 Jul 15. Leuk Lymphoma. 2020. PMID: 32667245
-
CD73 Promotes Chronic Lymphocytic Leukemia.Cancers (Basel). 2022 Jun 26;14(13):3130. doi: 10.3390/cancers14133130. Cancers (Basel). 2022. PMID: 35804900 Free PMC article.
-
Regulation of CLL survival by hypoxia-inducible factor and its target genes.FEBS Lett. 2012 Aug 31;586(18):2906-10. doi: 10.1016/j.febslet.2012.07.016. Epub 2012 Jul 24. FEBS Lett. 2012. PMID: 22841548 Review.
Cited by
-
Extracellular Vesicle Secretion by Leukemia Cells In Vivo Promotes CLL Progression by Hampering Antitumor T-cell Responses.Blood Cancer Discov. 2023 Jan 6;4(1):54-77. doi: 10.1158/2643-3230.BCD-22-0029. Blood Cancer Discov. 2023. PMID: 36108149 Free PMC article.
-
Hypoxic stress and hypoxia-inducible factors in leukemias.Front Oncol. 2022 Aug 18;12:973978. doi: 10.3389/fonc.2022.973978. eCollection 2022. Front Oncol. 2022. PMID: 36059690 Free PMC article. Review.
-
Indoleamine 2, 3-Dioxygenase 1 Mediates Survival Signals in Chronic Lymphocytic Leukemia via Kynurenine/Aryl Hydrocarbon Receptor-Mediated MCL1 Modulation.Front Immunol. 2022 Mar 18;13:832263. doi: 10.3389/fimmu.2022.832263. eCollection 2022. Front Immunol. 2022. PMID: 35371054 Free PMC article.
References
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

