The Role of Drosophila CtIP in Homology-Directed Repair of DNA Double-Strand Breaks
- PMID: 34573412
- PMCID: PMC8468788
- DOI: 10.3390/genes12091430
The Role of Drosophila CtIP in Homology-Directed Repair of DNA Double-Strand Breaks
Abstract
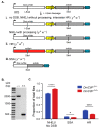
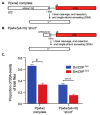
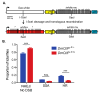
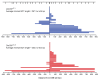
DNA double-strand breaks (DSBs) are a particularly genotoxic type of DNA damage that can result in chromosomal aberrations. Thus, proper repair of DSBs is essential to maintaining genome integrity. DSBs can be repaired by non-homologous end joining (NHEJ), where ends are processed before joining through ligation. Alternatively, DSBs can be repaired through homology-directed repair, either by homologous recombination (HR) or single-strand annealing (SSA). Both types of homology-directed repair are initiated by DNA end resection. In cultured human cells, the protein CtIP has been shown to play a role in DNA end resection through its interactions with CDK, BRCA1, DNA2, and the MRN complex. To elucidate the role of CtIP in a multicellular context, CRISPR/Cas9 genome editing was used to create a DmCtIPΔ allele in Drosophila melanogaster. Using the DSB repair reporter assay direct repeat of white (DR-white), a two-fold decrease in HR in DmCtIPΔ/Δ mutants was observed when compared to heterozygous controls. However, analysis of HR gene conversion tracts (GCTs) suggests DmCtIP plays a minimal role in determining GCT length. To assess the function of DmCtIP on both short (~550 bp) and long (~3.6 kb) end resection, modified homology-directed SSA repair assays were implemented, resulting in a two-fold decrease in SSA repair in both short and extensive end resection requirements in the DmCtIPΔ/Δ mutants compared to heterozygote controls. Through these analyses, we affirmed the importance of end resection on DSB repair pathway choice in multicellular systems, described the function of DmCtIP in short and extensive DNA end resection, and determined the impact of end resection on GCT length during HR.
Keywords: CtIP; Drosophila; double-strand break repair; end resection; homologous recombination; non-homologous end-joining; single-strand annealing.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
Similar articles
-
The epistatic relationship of Drosophila melanogaster CtIP and Rif1 in homology-directed repair of DNA double-strand breaks.G3 (Bethesda). 2024 Nov 6;14(11):jkae210. doi: 10.1093/g3journal/jkae210. G3 (Bethesda). 2024. PMID: 39397376 Free PMC article.
-
The impact of developmental stage, tissue type, and sex on DNA double-strand break repair in Drosophila melanogaster.PLoS Genet. 2024 Apr 29;20(4):e1011250. doi: 10.1371/journal.pgen.1011250. eCollection 2024 Apr. PLoS Genet. 2024. PMID: 38683763 Free PMC article.
-
Analysis of chromatid-break-repair detects a homologous recombination to non-homologous end-joining switch with increasing load of DNA double-strand breaks.Mutat Res Genet Toxicol Environ Mutagen. 2021 Jul;867:503372. doi: 10.1016/j.mrgentox.2021.503372. Epub 2021 Jun 12. Mutat Res Genet Toxicol Environ Mutagen. 2021. PMID: 34266628
-
Crosstalk between different DNA repair pathways for DNA double strand break repairs.Mutat Res Genet Toxicol Environ Mutagen. 2022 Jan;873:503438. doi: 10.1016/j.mrgentox.2021.503438. Epub 2021 Dec 15. Mutat Res Genet Toxicol Environ Mutagen. 2022. PMID: 35094810 Review.
-
DNA End Resection: Mechanism and Control.Annu Rev Genet. 2021 Nov 23;55:285-307. doi: 10.1146/annurev-genet-071719-020312. Annu Rev Genet. 2021. PMID: 34813349 Review.
Cited by
-
The epistatic relationship of Drosophila melanogaster CtIP and Rif1 in homology-directed repair of DNA double-strand breaks.G3 (Bethesda). 2024 Nov 6;14(11):jkae210. doi: 10.1093/g3journal/jkae210. G3 (Bethesda). 2024. PMID: 39397376 Free PMC article.
-
Transgene removal using an in cis programmed homing endonuclease via single-strand annealing in the mosquito Aedes aegypti.Commun Biol. 2024 May 29;7(1):660. doi: 10.1038/s42003-024-06348-6. Commun Biol. 2024. PMID: 38811748 Free PMC article.
-
Homing gene drives can transfer rapidly between Anopheles gambiae strains with minimal carryover of flanking sequences.Nat Commun. 2024 Aug 10;15(1):6846. doi: 10.1038/s41467-024-51225-9. Nat Commun. 2024. PMID: 39122734 Free PMC article.
-
DNA Damage Response Mechanisms in Model Systems.Genes (Basel). 2023 Jun 30;14(7):1385. doi: 10.3390/genes14071385. Genes (Basel). 2023. PMID: 37510290 Free PMC article.
-
Driving a protective allele of the mosquito FREP1 gene to combat malaria.Nature. 2025 Jul 23. doi: 10.1038/s41586-025-09283-6. Online ahead of print. Nature. 2025. PMID: 40702179
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

