Cross-neutralizing antibodies bind a SARS-CoV-2 cryptic site and resist circulating variants
- PMID: 34580306
- PMCID: PMC8476643
- DOI: 10.1038/s41467-021-25997-3
Cross-neutralizing antibodies bind a SARS-CoV-2 cryptic site and resist circulating variants
Abstract
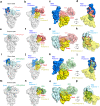
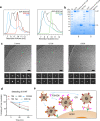
The emergence of numerous variants of SARS-CoV-2, the causative agent of COVID-19, has presented new challenges to the global efforts to control the COVID-19 pandemic. Here, we obtain two cross-neutralizing antibodies (7D6 and 6D6) that target Sarbecoviruses' receptor-binding domain (RBD) with sub-picomolar affinities and potently neutralize authentic SARS-CoV-2. Crystal structures show that both antibodies bind a cryptic site different from that recognized by existing antibodies and highly conserved across Sarbecovirus isolates. Binding of these two antibodies to the RBD clashes with the adjacent N-terminal domain and disrupts the viral spike. Both antibodies confer good resistance to mutations in the currently circulating SARS-CoV-2 variants. Thus, our results have direct relevance to public health as options for passive antibody therapeutics and even active prophylactics. They can also inform the design of pan-sarbecovirus vaccines.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




Similar articles
-
Escape from neutralizing antibodies by SARS-CoV-2 spike protein variants.Elife. 2020 Oct 28;9:e61312. doi: 10.7554/eLife.61312. Elife. 2020. PMID: 33112236 Free PMC article.
-
Cross-Neutralization of Emerging SARS-CoV-2 Variants of Concern by Antibodies Targeting Distinct Epitopes on Spike.mBio. 2021 Dec 21;12(6):e0297521. doi: 10.1128/mBio.02975-21. Epub 2021 Nov 16. mBio. 2021. PMID: 34781736 Free PMC article. Clinical Trial.
-
Computational design of a neutralizing antibody with picomolar binding affinity for all concerning SARS-CoV-2 variants.MAbs. 2022 Jan-Dec;14(1):2021601. doi: 10.1080/19420862.2021.2021601. MAbs. 2022. PMID: 35030983 Free PMC article.
-
Structural Analysis of Neutralizing Epitopes of the SARS-CoV-2 Spike to Guide Therapy and Vaccine Design Strategies.Viruses. 2021 Jan 19;13(1):134. doi: 10.3390/v13010134. Viruses. 2021. PMID: 33477902 Free PMC article. Review.
-
Analysis of the molecular mechanism of SARS-CoV-2 antibodies.Biochem Biophys Res Commun. 2021 Aug 20;566:45-52. doi: 10.1016/j.bbrc.2021.06.001. Epub 2021 Jun 5. Biochem Biophys Res Commun. 2021. PMID: 34116356 Free PMC article. Review.
Cited by
-
Two antibodies show broad, synergistic neutralization against SARS-CoV-2 variants by inducing conformational change within the RBD.Protein Cell. 2024 Feb 1;15(2):121-134. doi: 10.1093/procel/pwad040. Protein Cell. 2024. PMID: 37470320 Free PMC article.
-
Variations in cell-surface ACE2 levels alter direct binding of SARS-CoV-2 Spike protein and viral infectivity: Implications for measuring Spike protein interactions with animal ACE2 orthologs.bioRxiv [Preprint]. 2021 Oct 22:2021.10.21.465386. doi: 10.1101/2021.10.21.465386. bioRxiv. 2021. Update in: J Virol. 2022 Sep 14;96(17):e0025622. doi: 10.1128/jvi.00256-22. PMID: 34729559 Free PMC article. Updated. Preprint.
-
Nanobody repertoire generated against the spike protein of ancestral SARS-CoV-2 remains efficacious against the rapidly evolving virus.Elife. 2024 May 7;12:RP89423. doi: 10.7554/eLife.89423. Elife. 2024. PMID: 38712823 Free PMC article.
-
Evolving SARS-CoV-2 Vaccines: From Current Solutions to Broad-Spectrum Protection.Vaccines (Basel). 2025 Jun 12;13(6):635. doi: 10.3390/vaccines13060635. Vaccines (Basel). 2025. PMID: 40573967 Free PMC article. Review.
-
Omicron BA.1 breakthrough infection drives cross-variant neutralization and memory B cell formation against conserved epitopes.Sci Immunol. 2022 Sep 16;7(75):eabq2427. doi: 10.1126/sciimmunol.abq2427. Epub 2022 Sep 16. Sci Immunol. 2022. PMID: 35653438 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
- 82001756/National Natural Science Foundation of China (National Science Foundation of China)
- 82001756/National Natural Science Foundation of China (National Science Foundation of China)
- U1705283/National Natural Science Foundation of China (National Science Foundation of China)
- 81991491/National Natural Science Foundation of China (National Science Foundation of China)
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Miscellaneous

