The CXCL Family Contributes to Immunosuppressive Microenvironment in Gliomas and Assists in Gliomas Chemotherapy
- PMID: 34603309
- PMCID: PMC8482424
- DOI: 10.3389/fimmu.2021.731751
The CXCL Family Contributes to Immunosuppressive Microenvironment in Gliomas and Assists in Gliomas Chemotherapy
Abstract
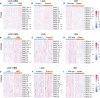
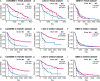
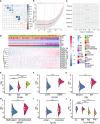
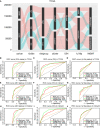
Gliomas are a type of malignant central nervous system tumor with poor prognosis. Molecular biomarkers of gliomas can predict glioma patient's clinical outcome, but their limitations are also emerging. C-X-C motif chemokine ligand family plays a critical role in shaping tumor immune landscape and modulating tumor progression, but its role in gliomas is elusive. In this work, samples of TCGA were treated as the training cohort, and as for validation cohort, two CGGA datasets, four datasets from GEO database, and our own clinical samples were enrolled. Consensus clustering analysis was first introduced to classify samples based on CXCL expression profile, and the support vector machine was applied to construct the cluster model in validation cohort based on training cohort. Next, the elastic net analysis was applied to calculate the risk score of each sample based on CXCL expression. High-risk samples associated with more malignant clinical features, worse survival outcome, and more complicated immune landscape than low-risk samples. Besides, higher immune checkpoint gene expression was also noticed in high-risk samples, suggesting CXCL may participate in tumor evasion from immune surveillance. Notably, high-risk samples also manifested higher chemotherapy resistance than low-risk samples. Therefore, we predicted potential compounds that target high-risk samples. Two novel drugs, LCL-161 and ADZ5582, were firstly identified as gliomas' potential compounds, and five compounds from PubChem database were filtered out. Taken together, we constructed a prognostic model based on CXCL expression, and predicted that CXCL may affect tumor progression by modulating tumor immune landscape and tumor immune escape. Novel potential compounds were also proposed, which may improve malignant glioma prognosis.
Keywords: CXCL; chemotherapy; gliomas; immune checkpoint genes; immunosuppressive.
Copyright © 2021 Wang, Liu, Mo, Zhang, Dai, Zhang, Ye, Cao, Liu and Cheng.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
Ferroptosis-related gene signature correlates with the tumor immune features and predicts the prognosis of glioma patients.Biosci Rep. 2021 Dec 22;41(12):BSR20211640. doi: 10.1042/BSR20211640. Biosci Rep. 2021. PMID: 34726238 Free PMC article.
-
Comprehensive exploration of tumor mutational burden and immune infiltration in diffuse glioma.Int Immunopharmacol. 2021 Jul;96:107610. doi: 10.1016/j.intimp.2021.107610. Epub 2021 Apr 10. Int Immunopharmacol. 2021. PMID: 33848908
-
Hypoxia-Related lncRNA Correlates With Prognosis and Immune Microenvironment in Lower-Grade Glioma.Front Immunol. 2021 Sep 30;12:731048. doi: 10.3389/fimmu.2021.731048. eCollection 2021. Front Immunol. 2021. PMID: 34659218 Free PMC article.
-
Epigenetic reprogramming and antitumor immune responses in gliomas: a systematic review.Med Oncol. 2025 May 16;42(6):213. doi: 10.1007/s12032-025-02760-y. Med Oncol. 2025. PMID: 40380049 Review.
-
Immune microenvironment of gliomas.Lab Invest. 2017 May;97(5):498-518. doi: 10.1038/labinvest.2017.19. Epub 2017 Mar 13. Lab Invest. 2017. PMID: 28287634 Review.
Cited by
-
Limitations of nomogram models in predicting survival outcomes for glioma patients.Front Immunol. 2025 Mar 18;16:1547506. doi: 10.3389/fimmu.2025.1547506. eCollection 2025. Front Immunol. 2025. PMID: 40170838 Free PMC article.
-
A Lamin Family-Based Signature Predicts Prognosis and Immunotherapy Response in Hepatocellular Carcinoma.J Immunol Res. 2022 Nov 10;2022:4983532. doi: 10.1155/2022/4983532. eCollection 2022. J Immunol Res. 2022. PMID: 36405011 Free PMC article.
-
Identification of prognostic risk score of disulfidptosis-related genes and molecular subtypes in glioma.Biochem Biophys Rep. 2023 Dec 17;37:101605. doi: 10.1016/j.bbrep.2023.101605. eCollection 2024 Mar. Biochem Biophys Rep. 2023. PMID: 38188362 Free PMC article.
-
CHRM3 is a novel prognostic factor of poor prognosis and promotes glioblastoma progression via activation of oncogenic invasive growth factors.Oncol Res. 2023 Sep 15;31(6):917-927. doi: 10.32604/or.2023.030425. eCollection 2023. Oncol Res. 2023. PMID: 37744266 Free PMC article.
-
Comprehensive analysis of CXCL10 and MIP-3a reveals their potential clinical application in hepatocellular carcinoma.Transl Oncol. 2024 Oct;48:102071. doi: 10.1016/j.tranon.2024.102071. Epub 2024 Aug 3. Transl Oncol. 2024. PMID: 39098213 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical

