Enzymology and Regulation of δ1-Pyrroline-5-Carboxylate Synthetase 2 From Rice
- PMID: 34603346
- PMCID: PMC8480329
- DOI: 10.3389/fpls.2021.672702
Enzymology and Regulation of δ1-Pyrroline-5-Carboxylate Synthetase 2 From Rice
Abstract
Under several stress conditions, such as excess salt and drought, many plants accumulate proline inside the cell, which is believed to help counteracting the adverse effects of low water potential. This increase mainly relies upon transcriptional induction of δ1-pyrroline-5-carboxylate synthetase (P5CS), the enzyme that catalyzes the first two steps in proline biosynthesis from glutamate. P5CS mediates both the phosphorylation of glutamate and the reduction of γ-glutamylphosphate to glutamate-5-semialdehyde, which spontaneously cyclizes to δ1-pyrroline-5-carboxylate (P5C). In most higher plants, two isoforms of P5CS have been found, one constitutively expressed to satisfy proline demand for protein synthesis, the other stress-induced. Despite the number of papers to investigate the regulation of P5CS at the transcriptional level, to date, the properties of the enzyme have been only poorly studied. As a consequence, the descriptions of post-translational regulatory mechanisms have largely been limited to feedback-inhibition by proline. Here, we report cloning and heterologous expression of P5CS2 from Oryza sativa. The protein has been fully characterized from a functional point of view, using an assay method that allows following the physiological reaction of the enzyme. Kinetic analyses show that the activity is subjected to a wide array of regulatory mechanisms, ranging from product inhibition to feedback inhibition by proline and other amino acids. These findings confirm long-hypothesized influences of both, the redox status of the cell and nitrogen availability, on proline biosynthesis.
Keywords: NADPH/NADP+ ratio; enzyme regulation; product inhibition; proline biosynthesis; redox status; substrate affinity.
Copyright © 2021 Sabbioni, Funck and Forlani.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
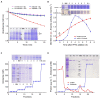
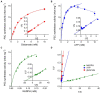
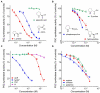
Figures


Similar articles
-
Removal of feedback inhibition of delta 1-pyrroline-5-carboxylate synthetase, a bifunctional enzyme catalyzing the first two steps of proline biosynthesis in plants.J Biol Chem. 1995 Sep 1;270(35):20491-6. doi: 10.1074/jbc.270.35.20491. J Biol Chem. 1995. PMID: 7657626
-
A complex array of factors regulate the activity of Arabidopsis thaliana δ1 -pyrroline-5-carboxylate synthetase isoenzymes to ensure their specific role in plant cell metabolism.Plant Cell Environ. 2024 Apr;47(4):1348-1362. doi: 10.1111/pce.14817. Epub 2024 Jan 15. Plant Cell Environ. 2024. PMID: 38223941
-
A bifunctional enzyme (delta 1-pyrroline-5-carboxylate synthetase) catalyzes the first two steps in proline biosynthesis in plants.Proc Natl Acad Sci U S A. 1992 Oct 1;89(19):9354-8. doi: 10.1073/pnas.89.19.9354. Proc Natl Acad Sci U S A. 1992. PMID: 1384052 Free PMC article.
-
P5C as an Interface of Proline Interconvertible Amino Acids and Its Role in Regulation of Cell Survival and Apoptosis.Int J Mol Sci. 2021 Oct 29;22(21):11763. doi: 10.3390/ijms222111763. Int J Mol Sci. 2021. PMID: 34769188 Free PMC article. Review.
-
Regulation of levels of proline as an osmolyte in plants under water stress.Plant Cell Physiol. 1997 Oct;38(10):1095-102. doi: 10.1093/oxfordjournals.pcp.a029093. Plant Cell Physiol. 1997. PMID: 9399433 Review.
Cited by
-
The Emerging Role of Proline in the Establishment and Functioning of Legume-Rhizobium Symbiosis.Front Plant Sci. 2022 May 27;13:888769. doi: 10.3389/fpls.2022.888769. eCollection 2022. Front Plant Sci. 2022. PMID: 35712558 Free PMC article. Review.
-
Integrative physiological, metabolomic, and transcriptomic analysis reveals the drought responses of two apple rootstock cultivars.BMC Plant Biol. 2024 Mar 26;24(1):219. doi: 10.1186/s12870-024-04902-2. BMC Plant Biol. 2024. PMID: 38532379 Free PMC article.
-
Integrative transcriptomic and metabolomic analysis elucidates the vital pathways underlying the differences in salt stress responses between two chickpea (Cicer arietinum L.) varieties.BMC Plant Biol. 2025 Jul 12;25(1):903. doi: 10.1186/s12870-025-06910-2. BMC Plant Biol. 2025. PMID: 40646445 Free PMC article.
-
Proline Affects Flowering Time in Arabidopsis by Modulating FLC Expression: A Clue of Epigenetic Regulation?Plants (Basel). 2022 Sep 8;11(18):2348. doi: 10.3390/plants11182348. Plants (Basel). 2022. PMID: 36145748 Free PMC article.
-
Deciphering molecular regulation of reactive oxygen species (ROS) and reactive nitrogen species (RNS) signalling networks in Oryza genus amid environmental stress.Plant Cell Rep. 2024 Jul 1;43(7):185. doi: 10.1007/s00299-024-03264-1. Plant Cell Rep. 2024. PMID: 38951279 Review.
References
-
- Ann-Brittedfast T. N., Ericsson A., Nordén L.-G. (1994). Accumulation of amino acids in some boreal forest plants in response to increased nitrogen availability. New Phytol. 126, 137–143. doi: 10.1111/j.1469-8137.1994.tb07539.x - DOI
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

