Structural and catalytic characterization of Blastochloris viridis and Pseudomonas aeruginosa homospermidine synthases supports the essential role of cation-π interaction
- PMID: 34605434
- PMCID: PMC8489232
- DOI: 10.1107/S2059798321008937
Structural and catalytic characterization of Blastochloris viridis and Pseudomonas aeruginosa homospermidine synthases supports the essential role of cation-π interaction
Abstract
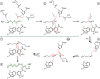
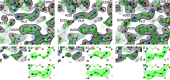
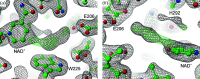
Polyamines influence medically relevant processes in the opportunistic pathogen Pseudomonas aeruginosa, including virulence, biofilm formation and susceptibility to antibiotics. Although homospermidine synthase (HSS) is part of the polyamine metabolism in various strains of P. aeruginosa, neither its role nor its structure has been examined so far. The reaction mechanism of the nicotinamide adenine dinucleotide (NAD+)-dependent bacterial HSS has previously been characterized based on crystal structures of Blastochloris viridis HSS (BvHSS). This study presents the crystal structure of P. aeruginosa HSS (PaHSS) in complex with its substrate putrescine. A high structural similarity between PaHSS and BvHSS with conservation of the catalytically relevant residues is demonstrated, qualifying BvHSS as a model for mechanistic studies of PaHSS. Following this strategy, crystal structures of single-residue variants of BvHSS are presented together with activity assays of PaHSS, BvHSS and BvHSS variants. For efficient homospermidine production, acidic residues are required at the entrance to the binding pocket (`ionic slide') and near the active site (`inner amino site') to attract and bind the substrate putrescine via salt bridges. The tryptophan residue at the active site stabilizes cationic reaction components by cation-π interaction, as inferred from the interaction geometry between putrescine and the indole ring plane. Exchange of this tryptophan for other amino acids suggests a distinct catalytic requirement for an aromatic interaction partner with a highly negative electrostatic potential. These findings substantiate the structural and mechanistic knowledge on bacterial HSS, a potential target for antibiotic design.
Keywords: Blastochloris viridis; NAD; Pseudomonas aeruginosa; Rossmann fold; cation–π interactions; homospermidine synthases; polyamine metabolism; putrescine; transferases.
open access.
Figures
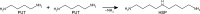





Similar articles
-
Rational engineering of homospermidine synthase for enhanced catalytic efficiency toward spermidine synthesis.Synth Syst Biotechnol. 2024 Apr 19;9(3):549-557. doi: 10.1016/j.synbio.2024.04.012. eCollection 2024 Sep. Synth Syst Biotechnol. 2024. PMID: 38699566 Free PMC article.
-
Comprehensive Structural Characterization of the Bacterial Homospermidine Synthase-an Essential Enzyme of the Polyamine Metabolism.Sci Rep. 2016 Jan 18;6:19501. doi: 10.1038/srep19501. Sci Rep. 2016. PMID: 26776105 Free PMC article.
-
Insights into the Unusual Activity of a Novel Homospermidine Synthase with a Promising Application to Produce Spermidine.J Agric Food Chem. 2023 Sep 6;71(35):13024-13034. doi: 10.1021/acs.jafc.3c03037. Epub 2023 Aug 25. J Agric Food Chem. 2023. PMID: 37622688
-
Structural basis of substrate binding specificity revealed by the crystal structures of polyamine receptors SpuD and SpuE from Pseudomonas aeruginosa.J Mol Biol. 2012 Mar 9;416(5):697-712. doi: 10.1016/j.jmb.2012.01.010. Epub 2012 Jan 28. J Mol Biol. 2012. PMID: 22300763
-
Evolution and multifarious horizontal transfer of an alternative biosynthetic pathway for the alternative polyamine sym-homospermidine.J Biol Chem. 2010 May 7;285(19):14711-23. doi: 10.1074/jbc.M110.107219. Epub 2010 Mar 1. J Biol Chem. 2010. PMID: 20194510 Free PMC article.
Cited by
-
Rational engineering of homospermidine synthase for enhanced catalytic efficiency toward spermidine synthesis.Synth Syst Biotechnol. 2024 Apr 19;9(3):549-557. doi: 10.1016/j.synbio.2024.04.012. eCollection 2024 Sep. Synth Syst Biotechnol. 2024. PMID: 38699566 Free PMC article.
References
-
- Anantharajah, A., Mingeot-Leclercq, M.-P. & Van Bambeke, F. (2016). Trends Pharmacol. Sci. 37, 734–749. - PubMed
-
- Becker, N. N. & Roberts, J. D. (1984). Biochemistry, 23, 3336–3340. - PubMed
-
- Benach, J., Atrian, S., Gonzàlez-Duarte, R. & Ladenstein, R. (1999). J. Mol. Biol. 289, 335–355. - PubMed
-
- Böttcher, F., Ober, D. & Hartmann, T. (1994). Can. J. Chem. 72, 80–85.
MeSH terms
Substances
Supplementary concepts
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

