Para-Aminobenzoic Acid, Calcium, and c-di-GMP Induce Formation of Cohesive, Syp-Polysaccharide-Dependent Biofilms in Vibrio fischeri
- PMID: 34607467
- PMCID: PMC8546588
- DOI: 10.1128/mBio.02034-21
Para-Aminobenzoic Acid, Calcium, and c-di-GMP Induce Formation of Cohesive, Syp-Polysaccharide-Dependent Biofilms in Vibrio fischeri
Abstract
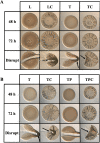
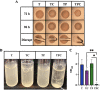
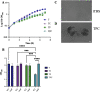
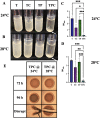
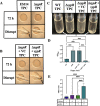
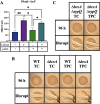
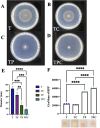
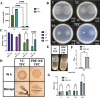
The marine bacterium Vibrio fischeri efficiently colonizes its symbiotic squid host, Euprymna scolopes, by producing a transient biofilm dependent on the symbiosis polysaccharide (SYP). In vitro, however, wild-type strain ES114 fails to form SYP-dependent biofilms. Instead, genetically engineered strains, such as those lacking the negative regulator BinK, have been developed to study this phenomenon. Historically, V. fischeri has been grown using LBS, a complex medium containing tryptone and yeast extract; supplementation with calcium is required to induce biofilm formation by a binK mutant. Here, through our discovery that yeast extract inhibits biofilm formation, we uncover signals and underlying mechanisms that control V. fischeri biofilm formation. In contrast to its inability to form a biofilm on unsupplemented LBS, a binK mutant formed cohesive, SYP-dependent colony biofilms on tTBS, modified LBS that lacks yeast extract. Moreover, wild-type strain ES114 became proficient to form cohesive, SYP-dependent biofilms when grown in tTBS supplemented with both calcium and the vitamin para-aminobenzoic acid (pABA); neither molecule alone was sufficient, indicating that this phenotype relies on coordinating two cues. pABA/calcium supplementation also inhibited bacterial motility. Consistent with these phenotypes, cells grown in tTBS with pABA/calcium were enriched in transcripts for biofilm-related genes and predicted diguanylate cyclases, which produce the second messenger cyclic-di-GMP (c-di-GMP). They also exhibited elevated levels of c-di-GMP, which was required for the observed phenotypes, as phosphodiesterase overproduction abrogated biofilm formation and partially rescued motility. This work thus provides insight into conditions, signals, and processes that promote biofilm formation by V. fischeri. IMPORTANCE Bacteria integrate environmental signals to regulate gene expression and protein production to adapt to their surroundings. One such behavioral adaptation is the formation of a biofilm, which can promote adherence and colonization and provide protection against antimicrobials. Identifying signals that trigger biofilm formation and the underlying mechanism(s) of action remain important and challenging areas of investigation. Here, we determined that yeast extract, commonly used for growth of bacteria in laboratory culture, inhibits biofilm formation by Vibrio fischeri, a model bacterium used for investigating host-relevant biofilm formation. Omitting yeast extract from the growth medium led to the identification of an unusual signal, the vitamin para-aminobenzoic acid (pABA), that when added together with calcium could induce biofilm formation. pABA increased the concentrations of the second messenger, c-di-GMP, which was necessary but not sufficient to induce biofilm formation. This work thus advances our understanding of signals and signal integration controlling bacterial biofilm formation.
Keywords: Vibrio fischeri; biofilms; c-di-GMP; calcium signaling; cyclic nucleotides; pABA; signal transduction.
Figures

Similar articles
-
A single point mutation is sufficient to drive syp-dependent biofilm formation and promote colonization by Vibrio fischeri.J Bacteriol. 2025 Aug 21;207(8):e0013125. doi: 10.1128/jb.00131-25. Epub 2025 Jul 14. J Bacteriol. 2025. PMID: 40657914 Free PMC article.
-
High Levels of Cyclic Diguanylate Interfere with Beneficial Bacterial Colonization.mBio. 2022 Aug 30;13(4):e0167122. doi: 10.1128/mbio.01671-22. Epub 2022 Aug 2. mBio. 2022. PMID: 35916402 Free PMC article.
-
Genetic Analysis Reveals a Requirement for the Hybrid Sensor Kinase RscS in para-Aminobenzoic Acid/Calcium-Induced Biofilm Formation by Vibrio fischeri.J Bacteriol. 2023 Jul 25;205(7):e0007523. doi: 10.1128/jb.00075-23. Epub 2023 Jun 12. J Bacteriol. 2023. PMID: 37306594 Free PMC article.
-
Control of biofilm formation and colonization in Vibrio fischeri: a role for partner switching?Environ Microbiol. 2010 Aug;12(8):2051-9. doi: 10.1111/j.1462-2920.2010.02269.x. Epub 2010 Jun 9. Environ Microbiol. 2010. PMID: 21966901 Free PMC article. Review.
-
Vibrio fischeri: a model for host-associated biofilm formation.J Bacteriol. 2024 Feb 22;206(2):e0037023. doi: 10.1128/jb.00370-23. Epub 2024 Jan 25. J Bacteriol. 2024. PMID: 38270381 Free PMC article. Review.
Cited by
-
A single point mutation is sufficient to drive syp-dependent biofilm formation and promote colonization by Vibrio fischeri.J Bacteriol. 2025 Aug 21;207(8):e0013125. doi: 10.1128/jb.00131-25. Epub 2025 Jul 14. J Bacteriol. 2025. PMID: 40657914 Free PMC article.
-
Transcriptional pathways across colony biofilm models in the symbiont Vibrio fischeri.mSystems. 2024 Jan 23;9(1):e0081523. doi: 10.1128/msystems.00815-23. Epub 2023 Dec 21. mSystems. 2024. PMID: 38126773 Free PMC article.
-
Characterization of bacteria colonizing the mucosal layer of the gastrointestinal tract of Atlantic salmon farmed in a warm water region.Front Microbiol. 2025 Jul 23;16:1564052. doi: 10.3389/fmicb.2025.1564052. eCollection 2025. Front Microbiol. 2025. PMID: 40771687 Free PMC article.
-
Genetic and biochemical analyses reveal direct interactions between LitR and genes important for Vibrio fischeri physiology, including biofilm production.J Bacteriol. 2025 Aug 21;207(8):e0004225. doi: 10.1128/jb.00042-25. Epub 2025 Aug 1. J Bacteriol. 2025. PMID: 40748066 Free PMC article.
-
High Levels of Cyclic Diguanylate Interfere with Beneficial Bacterial Colonization.mBio. 2022 Aug 30;13(4):e0167122. doi: 10.1128/mbio.01671-22. Epub 2022 Aug 2. mBio. 2022. PMID: 35916402 Free PMC article.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

