Spatial rank-based multifactor dimensionality reduction to detect gene-gene interactions for multivariate phenotypes
- PMID: 34607566
- PMCID: PMC8489107
- DOI: 10.1186/s12859-021-04395-y
Spatial rank-based multifactor dimensionality reduction to detect gene-gene interactions for multivariate phenotypes
Abstract
Background: Identifying interaction effects between genes is one of the main tasks of genome-wide association studies aiming to shed light on the biological mechanisms underlying complex diseases. Multifactor dimensionality reduction (MDR) is a popular approach for detecting gene-gene interactions that has been extended in various forms to handle binary and continuous phenotypes. However, only few multivariate MDR methods are available for multiple related phenotypes. Current approaches use Hotelling's T2 statistic to evaluate interaction models, but it is well known that Hotelling's T2 statistic is highly sensitive to heavily skewed distributions and outliers.
Results: We propose a robust approach based on nonparametric statistics such as spatial signs and ranks. The new multivariate rank-based MDR (MR-MDR) is mainly suitable for analyzing multiple continuous phenotypes and is less sensitive to skewed distributions and outliers. MR-MDR utilizes fuzzy k-means clustering and classifies multi-locus genotypes into two groups. Then, MR-MDR calculates a spatial rank-sum statistic as an evaluation measure and selects the best interaction model with the largest statistic. Our novel idea lies in adopting nonparametric statistics as an evaluation measure for robust inference. We adopt tenfold cross-validation to avoid overfitting. Intensive simulation studies were conducted to compare the performance of MR-MDR with current methods. Application of MR-MDR to a real dataset from a Korean genome-wide association study demonstrated that it successfully identified genetic interactions associated with four phenotypes related to kidney function. The R code for conducting MR-MDR is available at https://github.com/statpark/MR-MDR .
Conclusions: Intensive simulation studies comparing MR-MDR with several current methods showed that the performance of MR-MDR was outstanding for skewed distributions. Additionally, for symmetric distributions, MR-MDR showed comparable power. Therefore, we conclude that MR-MDR is a useful multivariate non-parametric approach that can be used regardless of the phenotype distribution, the correlations between phenotypes, and sample size.
Keywords: Fuzzy clustering; Gene–gene interaction; Multifactor dimensionality reduction; Spatial rank statistic.
© 2021. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
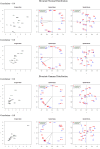
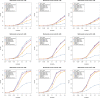
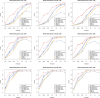
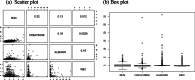
Figures

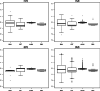
Similar articles
-
Multivariate Cluster-Based Multifactor Dimensionality Reduction to Identify Genetic Interactions for Multiple Quantitative Phenotypes.Biomed Res Int. 2019 Jul 11;2019:4578983. doi: 10.1155/2019/4578983. eCollection 2019. Biomed Res Int. 2019. PMID: 31380425 Free PMC article.
-
A unified model based multifactor dimensionality reduction framework for detecting gene-gene interactions.Bioinformatics. 2016 Sep 1;32(17):i605-i610. doi: 10.1093/bioinformatics/btw424. Bioinformatics. 2016. PMID: 27587680
-
Multivariate Quantitative Multifactor Dimensionality Reduction for Detecting Gene-Gene Interactions.Hum Hered. 2015;79(3-4):168-81. doi: 10.1159/000377723. Epub 2015 Jul 28. Hum Hered. 2015. PMID: 26201702
-
Epistasis, complexity, and multifactor dimensionality reduction.Methods Mol Biol. 2013;1019:465-77. doi: 10.1007/978-1-62703-447-0_22. Methods Mol Biol. 2013. PMID: 23756906 Review.
-
A roadmap to multifactor dimensionality reduction methods.Brief Bioinform. 2016 Mar;17(2):293-308. doi: 10.1093/bib/bbv038. Epub 2015 Jun 24. Brief Bioinform. 2016. PMID: 26108231 Free PMC article. Review.
Cited by
-
Melatonin Receptor 1B Genetic Variants on Susceptibility to Gestational Diabetes Mellitus: A Hospital-Based Case-Control Study in Wuhan, Central China.Diabetes Metab Syndr Obes. 2022 Apr 20;15:1207-1216. doi: 10.2147/DMSO.S345036. eCollection 2022. Diabetes Metab Syndr Obes. 2022. PMID: 35480849 Free PMC article.
-
Overview of frequent pattern mining.Genomics Inform. 2022 Dec;20(4):e39. doi: 10.5808/gi.22074. Epub 2022 Dec 30. Genomics Inform. 2022. PMID: 36617647 Free PMC article.
-
Predicting susceptibility to COVID-19 infection in patients on maintenance hemodialysis by cross-coupling soluble ACE2 concentration with lymphocyte count: an algorithmic approach.Front Med (Lausanne). 2024 Oct 30;11:1444719. doi: 10.3389/fmed.2024.1444719. eCollection 2024. Front Med (Lausanne). 2024. PMID: 39540040 Free PMC article.
-
A Hierarchical Error Correction Strategy for Text DNA Storage.Interdiscip Sci. 2022 Mar;14(1):141-150. doi: 10.1007/s12539-021-00476-x. Epub 2021 Aug 31. Interdiscip Sci. 2022. PMID: 34463928
References
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

