TSSK3, a novel target for male contraception, is required for spermiogenesis
- PMID: 34623009
- PMCID: PMC8961454
- DOI: 10.1002/mrd.23539
TSSK3, a novel target for male contraception, is required for spermiogenesis
Abstract
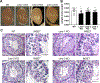
We have previously shown that members of the family of testis-specific serine/threonine kinases (TSSKs) are post-meiotically expressed in testicular germ cells and in mature sperm in mammals. The restricted post-meiotic expression of TSSKs as well as the importance of phosphorylation in signaling processes strongly suggest that TSSKs have an important role in germ cell differentiation and/or sperm function. This prediction has been supported by the reported sterile phenotype of the TSSK6 knock-out (KO) mice and of the double TSSK1/TSSK2 KO. The aim of this study was to develop KO mouse models of TSSK3 and to validate this kinase as a target for the development of a male contraceptive. We used CRISPR/Cas9 technology to generate the TSSK3 KO allele on B6D2F1 background mice. Male heterozygous pups were used to establish three independent TSSK3 KO lines. After natural mating of TSSK3 KO males, females that presented a plug (indicative of mating) were monitored for the following 24 days and no pregnancies or pups were found. Sperm numbers were drastically reduced in all three KO lines and, remarkably, round spermatids were detected in the cauda epididymis of KO mice. From the small population of sperm recovered, severe morphology defects were detected. Our results indicate an essential role of TSSK3 in spermiogenesis and support this kinase as a suitable candidate for the development of novel nonhormonal male contraceptives.
Keywords: TSSK3; evolution; fertilization; intronless gene; kinases; nonhormonal male contraceptive; sperm; spermatogenesis; testis-specific.
© 2021 Wiley Periodicals LLC.
Conflict of interest statement
Conflict of interest disclosure
Dr. Visconti and Dr. Salicioni own equity interest in Sperm Capacitation Technologies Inc., a company with goals in improving assisted reproductive technologies. The other authors have not conflict of interest to declare.
Figures


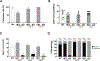


Similar articles
-
Testis-specific serine kinase 3 is required for sperm morphogenesis and male fertility.Andrology. 2023 Jul;11(5):826-839. doi: 10.1111/andr.13314. Epub 2022 Nov 16. Andrology. 2023. PMID: 36306217 Free PMC article.
-
Identification of TSSK1 and TSSK2 as Novel Targets for Male Contraception.Biomolecules. 2025 Apr 18;15(4):601. doi: 10.3390/biom15040601. Biomolecules. 2025. PMID: 40305308 Free PMC article.
-
Testis-specific serine kinase protein family in male fertility and as targets for non-hormonal male contraception†.Biol Reprod. 2020 Aug 4;103(2):264-274. doi: 10.1093/biolre/ioaa064. Biol Reprod. 2020. PMID: 32337545 Free PMC article. Review.
-
Expression and localization of five members of the testis-specific serine kinase (Tssk) family in mouse and human sperm and testis.Mol Hum Reprod. 2011 Jan;17(1):42-56. doi: 10.1093/molehr/gaq071. Epub 2010 Aug 20. Mol Hum Reprod. 2011. PMID: 20729278 Free PMC article.
-
Validation of a testis specific serine/threonine kinase [TSSK] family and the substrate of TSSK1 & 2, TSKS, as contraceptive targets.Soc Reprod Fertil Suppl. 2007;63:87-101. Soc Reprod Fertil Suppl. 2007. PMID: 17566264 Review.
Cited by
-
Advances in Male Contraception: When Will the Novel Male Contraception be Available?World J Mens Health. 2024 Jul;42(3):487-501. doi: 10.5534/wjmh.230118. Epub 2024 Jan 2. World J Mens Health. 2024. PMID: 38164023 Free PMC article. Review.
-
Characterization of sexual maturity-associated N6-methyladenosine in boar testes.BMC Genomics. 2024 May 7;25(1):447. doi: 10.1186/s12864-024-10343-w. BMC Genomics. 2024. PMID: 38714941 Free PMC article.
-
Testis-specific serine kinase 3 is required for sperm morphogenesis and male fertility.Andrology. 2023 Jul;11(5):826-839. doi: 10.1111/andr.13314. Epub 2022 Nov 16. Andrology. 2023. PMID: 36306217 Free PMC article.
-
Long COVID-19 autoantibodies and their potential effect on fertility.Front Immunol. 2025 May 27;16:1540341. doi: 10.3389/fimmu.2025.1540341. eCollection 2025. Front Immunol. 2025. PMID: 40496870 Free PMC article.
-
Identification of YBX2 and TSKS As STK33 Interacting Proteins in Testicular Germ Cells.Reprod Sci. 2025 Mar;32(3):769-782. doi: 10.1007/s43032-025-01798-7. Epub 2025 Feb 5. Reprod Sci. 2025. PMID: 39909973
References
-
- Bielke W, Blaschke RJ, Miescher GC, Zurcher G, Andres AC, and Ziemiecki A. (1994). Characterization of a novel murine testis-specific serine/threonine kinase. Gene 139, 235–239. - PubMed
-
- Boutet I, Moraga D, Marinovic L, Obreque J, and Chavez-Crooker P. (2008). Characterization of reproduction-specific genes in a marine bivalve mollusc: influence of maturation stage and sex on mRNA expression. Gene 407, 130–138. - PubMed
-
- Bucko-Justyna M, Lipinski L, Burgering BM, and Trzeciak L. (2005). Characterization of testis-specific serine-threonine kinase 3 and its activation by phosphoinositide-dependent kinase-1-dependent signalling. The FEBS journal 272, 6310–6323. - PubMed
-
- Chemes HE (2013). Ultrastructural analysis of testicular tissue and sperm by transmission and scanning electron microscopy. Methods in molecular biology 927, 321–348. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

