Differential Expression of the TLR4 Gene in Pan-Cancer and Its Related Mechanism
- PMID: 34631699
- PMCID: PMC8495169
- DOI: 10.3389/fcell.2021.700661
Differential Expression of the TLR4 Gene in Pan-Cancer and Its Related Mechanism
Abstract
Previous studies have revealed the relationship between toll-like receptor 4 (TLR4) polymorphisms and cancer susceptibility. However, the relationship between TLR4 and prognosis and immune cell infiltration in pan-cancer patients is still unclear. Through the Genotype-Tissue Expression (GTEx) and The Cancer Genome Atlas (TCGA) databases, the distinct expression of the TLR4 gene in 24 tumors and normal tissues was analyzed. Univariate Cox proportional hazards regression analysis was used to identify the cancer types whose TLR4 gene expression was related to prognosis. The relationship between TLR4 and tumor cell immune invasion was studied. Spearman's rank correlation coefficient was used to analyze the relationship among TLR4 and immune neoantigens, tumor mutation burden (TMB), microsatellite instability (MSI), DNA repair genes, and DNA methylation. Gene Set Enrichment Analysis (GSEA) was used to identify the tumor-related pathways that the TLR4 gene was highly expressed in; the expression of the TLR4 gene was verified with the Human Protein Atlas (HPA) database. Low expression of TLR4 was associated with an inferior prognosis in kidney renal clear cell carcinoma (KIRC), skin cutaneous melanoma (SKCM), and uterine corpus endometrial carcinoma (UCEC), while high expression was related to a poor prognosis in head and neck squamous cell carcinoma (HNSC), prostate adenocarcinoma (PRAD), stomach adenocarcinoma (STAD), and testicular germ cell tumor (TGCT). The expression of TLR4 was negatively correlated with the expression of B cells in STAD. The expression of TLR4 was positively correlated with the infiltration of B cells, CD4 and CD8 T cells, neutrophils, macrophages, and dendritic cells in STAD, KIRC, UCEC, TGCT, and SKCM. The expression of the TLR4 gene in KIRC, SKCM, STAD, TGCT, and UCEC was highly correlated with inducible T-cell costimulator (ICOS), cytotoxic T lymphocyte-associated molecule 4 (CTLA4), and CD28 immune checkpoints. Spearman's rank correlation coefficient showed that the expression of TLR4 gene was significantly correlated with TMB in STAD and UCEC and was prominently correlated with MSI in TGCT, STAD, and SKCM. The expression of the TLR4 gene was highly correlated with MLH1, MSH2, and MSH6 in KIRC, SKCM, and STAD. The expression of the TLR4 gene was remarkably correlated with the methyltransferases DNA methyltransferase 2 (DNMT2) and DNA methyltransferase 3-beta (DNMT3B) in SKCM and STAD. Enrichment analysis showed that TLR4 was highly expressed in the chemokine signaling pathway and the cell adhesion molecule and cytokine receptor interaction pathway. In summary, the expression of TLR4 is linked to the prognosis of KIRC, SKCM, STAD, TGCT, and UCEC patients and the level of immune infiltration of CD4, CD8 T cells, macrophages, neutrophils, and dendritic cells.
Keywords: bioinformatics; immune cell infiltration; pan-cancer; prognosis; toll-like receptor 4.
Copyright © 2021 Hu, Xu, Feng, Li, Hua and Xu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




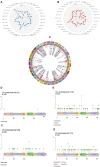



References
-
- Azimi F., Scolyer R. A., Rumcheva P., Moncrieff M., Murali R., McCarthy S. W., et al. (2012). Tumor-infiltrating lymphocyte grade is an independent predictor of sentinel lymph node status and survival in patients with cutaneous melanoma. J. Clin. Oncol. 30 2678–2683. 10.1200/jco.2011.37.8539 - DOI - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

