Generation and Staging of Human Retinal Organoids Based on Self-Formed Ectodermal Autonomous Multi-Zone System
- PMID: 34631711
- PMCID: PMC8493070
- DOI: 10.3389/fcell.2021.732382
Generation and Staging of Human Retinal Organoids Based on Self-Formed Ectodermal Autonomous Multi-Zone System
Abstract
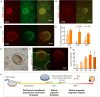
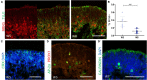
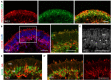
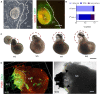
Methods for stem cell-derived, three-dimensional retinal organoids induction have been established and shown great potential for retinal development modeling and drug screening. Herein, we reported an exogenous-factors-free and robust method to generate retinal organoids based on "self-formed ectodermal autonomous multi-zone" (SEAM) system, a two-dimensional induction scheme that can synchronously generate multiple ocular cell lineages. Characterized by distinct morphological changes, the differentiation of the obtained retinal organoids could be staged into the early and late differentiation phases. During the early differentiation stage, retinal ganglion cells, cone photoreceptor cells (PRs), amacrine cells, and horizontal cells developed; whereas rod PRs, bipolar cells, and Müller glial cells were generated in the late differentiation phase, resembling early-phase and late-phase retinogenesis in vivo. Additionally, we modified the maintenance strategy for the retinal organoids and successfully promoted their long-term survival. Using 3D immunofluorescence image reconstruction and transmission electron microscopy, the substantial mature PRs with outer segment, inner segment and ribbon synapse were demonstrated. Besides, the retinal pigment epithelium (RPE) was induced with distinct boundary and the formation of ciliary margin was observed by co-suspending retina organoids with the zone containing RPE. The obtained RPE could be expanded and displayed similar marker expression, ultrastructural feature and functional phagocytosis to native RPE. Thus, this research described a simple and robust system which enabled generation of retina organoids with substantial mature PRs, RPE and the ciliary margin without the need of exogenous factors, providing a new platform for research of retinogenesis and retinal translational application.
Keywords: RPE; SEAM system; ciliary margin; human retinal organoid; photoreceptor cell; retinogenesis.
Copyright © 2021 Li, Chen, Ouyang, Ma, Sun, Luo, Chen and Liu.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures




Similar articles
-
One-stop assembly of adherent 3D retinal organoids from hiPSCs based on 3D-printed derived PDMS microwell platform.Biofabrication. 2023 Apr 11;15(3). doi: 10.1088/1758-5090/acc761. Biofabrication. 2023. PMID: 36963105
-
Accelerated photoreceptor differentiation of hiPSC-derived retinal organoids by contact co-culture with retinal pigment epithelium.Stem Cell Res. 2019 Aug;39:101491. doi: 10.1016/j.scr.2019.101491. Epub 2019 Jul 2. Stem Cell Res. 2019. PMID: 31326746
-
Decellularised extracellular matrix-derived peptides from neural retina and retinal pigment epithelium enhance the expression of synaptic markers and light responsiveness of human pluripotent stem cell derived retinal organoids.Biomaterials. 2019 Apr;199:63-75. doi: 10.1016/j.biomaterials.2019.01.028. Epub 2019 Jan 22. Biomaterials. 2019. PMID: 30738336
-
[Generation of Multiple Ocular Lineages from Human Pluripotent Stem Cells and Its Application to Regenerative Medicine].Yakugaku Zasshi. 2021;141(1):55-60. doi: 10.1248/yakushi.20-00177-4. Yakugaku Zasshi. 2021. PMID: 33390448 Review. Japanese.
-
Retinal organoids provide unique insights into molecular signatures of inherited retinal disease throughout retinogenesis.J Anat. 2023 Aug;243(2):186-203. doi: 10.1111/joa.13768. Epub 2022 Sep 29. J Anat. 2023. PMID: 36177499 Free PMC article. Review.
Cited by
-
A minimal-complexity light-sheet microscope maps network activity in 3D neuronal systems.Sci Rep. 2022 Nov 28;12(1):20420. doi: 10.1038/s41598-022-24350-y. Sci Rep. 2022. PMID: 36443413 Free PMC article.
-
The Analysis of Embryoid Body Formation and Its Role in Retinal Organoid Development.Int J Mol Sci. 2024 Jan 24;25(3):1444. doi: 10.3390/ijms25031444. Int J Mol Sci. 2024. PMID: 38338722 Free PMC article.
-
Single-cell transcriptomics reveals the molecular basis of human iPS cell differentiation into ectodermal ocular lineages.Commun Biol. 2024 Nov 12;7(1):1495. doi: 10.1038/s42003-024-07130-4. Commun Biol. 2024. PMID: 39532995 Free PMC article.
-
Rapid and efficient generation of mature retinal organoids derived from human pluripotent stem cells via optimized pharmacological modulation of Sonic hedgehog, activin A, and retinoic acid signal transduction.PLoS One. 2024 Aug 9;19(8):e0308743. doi: 10.1371/journal.pone.0308743. eCollection 2024. PLoS One. 2024. PMID: 39121095 Free PMC article.
-
Generating Retinas through Guided Pluripotent Stem Cell Differentiation and Direct Somatic Cell Reprogramming.Curr Stem Cell Res Ther. 2024;19(9):1251-1262. doi: 10.2174/011574888X255496230923164547. Curr Stem Cell Res Ther. 2024. PMID: 37807418 Review.
References
LinkOut - more resources
Full Text Sources

