Domain Organization of the UBX Domain Containing Protein 9 and Analysis of Its Interactions With the Homohexameric AAA + ATPase p97 (Valosin-Containing Protein)
- PMID: 34631722
- PMCID: PMC8495200
- DOI: 10.3389/fcell.2021.748860
Domain Organization of the UBX Domain Containing Protein 9 and Analysis of Its Interactions With the Homohexameric AAA + ATPase p97 (Valosin-Containing Protein)
Abstract
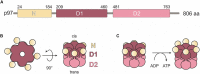
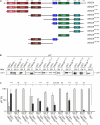
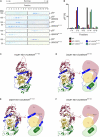
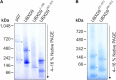
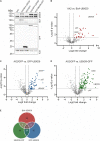
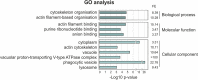
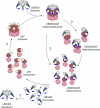
The abundant homohexameric AAA + ATPase p97 (also known as valosin-containing protein, VCP) is highly conserved from Dictyostelium discoideum to human and a pivotal factor of cellular protein homeostasis as it catalyzes the unfolding of proteins. Owing to its fundamental function in protein quality control pathways, it is regulated by more than 30 cofactors, including the UBXD protein family, whose members all carry an Ubiquitin Regulatory X (UBX) domain that enables binding to p97. One member of this latter protein family is the largely uncharacterized UBX domain containing protein 9 (UBXD9). Here, we analyzed protein-protein interactions of D. discoideum UBXD9 with p97 using a series of N- and C-terminal truncation constructs and probed the UBXD9 interactome in D. discoideum. Pull-down assays revealed that the UBX domain (amino acids 384-466) is necessary and sufficient for p97 interactions and that the N-terminal extension of the UBX domain, which folds into a β0-α- 1-α0 lariat structure, is required for the dissociation of p97 hexamers. Functionally, this finding is reflected by strongly reduced ATPase activity of p97 upon addition of full length UBXD9 or UBXD9261-573. Results from Blue Native PAGE as well as structural model prediction suggest that hexamers of UBXD9 or UBXD9261-573 interact with p97 hexamers and disrupt the p97 subunit interactions via insertion of a helical lariat structure, presumably by destabilizing the p97 D1:D1' intermolecular interface. We thus propose that UBXD9 regulates p97 activity in vivo by shifting the quaternary structure equilibrium from hexamers to monomers. Using three independent approaches, we further identified novel interaction partners of UBXD9, including glutamine synthetase type III as well as several actin-binding proteins. These findings suggest a role of UBXD9 in the organization of the actin cytoskeleton, and are in line with the hypothesized oligomerization-dependent mechanism of p97 regulation.
Keywords: ALS (Amyotrophic Lateral Sclerosis); ASPL; Dictyostelium discoideum; IBMPFD (Inclusion Body Myopathy associated with Paget disease of bone and Fronto temporal Dementia); PUX1); TUG; UBX domain containing protein 9 (UBXD9; hexamer disassembly; p97/VCP/CDC48/TER/VAT ATPase.
Copyright © 2021 Riehl, Rijal, Nitz, Clemen, Hofmann and Eichinger.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

References
-
- Abraham M. J., Murtola T., Schulz R., Páll S., Smith J. C., Hess B., et al. (2015). GROMACS: high performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1-2 19–25. 10.1016/j.softx.2015.06.001 - DOI
-
- Banchenko S., Arumughan A., Petrovic S., Schwefel D., Wanker E. E., Roske Y., et al. (2019). Common mode of remodeling AAA ATPases p97/CDC48 by their disassembling cofactors ASPL/PUX1. Structure 27 1830–1841.e3. - PubMed
LinkOut - more resources
Full Text Sources
Miscellaneous

