FLT4/VEGFR3 activates AMPK to coordinate glycometabolic reprogramming with autophagy and inflammasome activation for bacterial elimination
- PMID: 34632918
- PMCID: PMC9225223
- DOI: 10.1080/15548627.2021.1985338
FLT4/VEGFR3 activates AMPK to coordinate glycometabolic reprogramming with autophagy and inflammasome activation for bacterial elimination
Abstract
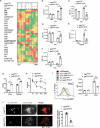
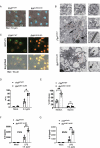
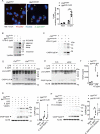
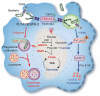
Macrophages rapidly undergo glycolytic reprogramming in response to macroautophagy/autophagy, inflammasome activation and pyroptosis for the clearance of bacteria. Identification the key molecules involved in these three events will provide critical potential therapeutic applications. Upon S. typhimurium infection, FLT4/VEGFR3 and its ligand VEGFC were inducibly expressed in macrophages, and FLT4 signaling inhibited CASP1 (caspase 1)-dependent inflammasome activation and pyroptosis but enhanced MAP1LC3/LC3 activation for elimination of the bacteria. Consistently, FLT4 mutants lacking the extracellular ligand-binding domain increased production of the proinflammatory metabolites such as succinate and lactate, and reduced antimicrobial metabolites including citrate and NAD(P)H in macrophages and liver upon infection. Mechanistically, FLT4 recruited AMP-activated protein kinase (AMPK) and phosphorylated Y247 and Y441/442 in the PRKAA/alpha subunit for AMPK activation. The AMPK agonist AICAR could rescue glycolytic reprogramming and inflammasome activation in macrophages expressing the mutant FLT4, which has potential translational application in patients carrying Flt4 mutations to prevent recurrent infections. Collectively, we have elucidated that the FLT4-AMPK module in macrophages coordinates glycolytic reprogramming, autophagy, inflammasome activation and pyroptosis to eliminate invading bacteria.Abbreviations: 3-MA: 3-methyladenine; AICAR: 5-aminoimidazole-4-carboxamide1-β-D-ribofuranoside; AMP: adenosine monophosphate; AMPK: AMP-activated protein kinase; ATP: adenosine triphosphate; BMDM: bone marrow-derived macrophage; CASP1: caspase 1; CFUs: colony-forming units; FLT4/VEGFR3: FMS-like tyrosine kinase 4; GFP: green fluorescent protein; LDH: lactate dehydrogenase; LPS: lipopolysaccharide; MAP1LC3/LC3: microtubule-associated protein 1 light chain 3; PEM: peritoneal exudate macrophage; PRKAA1/AMPKα1: protein kinase, AMP-activated, alpha 1 catalytic subunit; PYCARD/ASC: PYD and CARD domain containing; ROS: reactive oxygen species; SQSTM1/p62: sequestosome 1; TLR4: toll-like receptor 4; ULK1: unc-51 like autophagy activating kinase 1; VEGFC: vascular endothelial growth factor C; WT: wild type.
Keywords: AMPK; FLT4/VEGFR3; glycolytic reprogramming; inflammasome; pyroptosis.
Conflict of interest statement
The authors declare no competing financial interests.
Figures



References
-
- Kawai T, Akira S.. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat Immunol. 2010. May;11(5):373–384. - PubMed
-
- Van den Bossche J, O’Neill LA, Menon D. Macrophage immunometabolism: where are we (going)? Trends Immunol. 2017. June;38(6):395–406. - PubMed
-
- O’Neill LA. A critical role for citrate metabolism in LPS signalling. Biochem J. 2011. Sep;438(3):e5–6. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials
Miscellaneous
