Reductive enzymatic dynamic kinetic resolution affording 115 g/L (S)-2-phenylpropanol
- PMID: 34635076
- PMCID: PMC8507385
- DOI: 10.1186/s12896-021-00715-5
Reductive enzymatic dynamic kinetic resolution affording 115 g/L (S)-2-phenylpropanol
Abstract
Background: Published biocatalytic routes for accessing enantiopure 2-phenylpropanol using oxidoreductases afforded maximal product titers of only 80 mM. Enzyme deactivation was identified as the major limitation and was attributed to adduct formation of the aldehyde substrate with amino acid residues of the reductase.
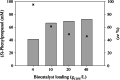
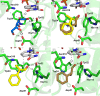
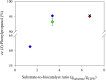
Results: A single point mutant of Candida tenuis xylose reductase (CtXR D51A) with very high catalytic efficiency (43·103 s-1 M-1) for (S)-2-phenylpropanal was found. The enzyme showed high enantioselectivity for the (S)-enantiomer but was deactivated by 0.5 mM substrate within 2 h. A whole-cell biocatalyst expressing the engineered reductase and a yeast formate dehydrogenase for NADH-recycling provided substantial stabilization of the reductase. The relatively slow in situ racemization of 2-phenylpropanal and the still limited biocatalyst stability required a subtle adjustment of the substrate-to-catalyst ratio. A value of 3.4 gsubstrate/gcell-dry-weight was selected as a suitable compromise between product ee and the conversion ratio. A catalyst loading of 40 gcell-dry-weight was used to convert 1 M racemic 2-phenylpropanal into 843 mM (115 g/L) (S)-phenylpropanol with 93.1% ee.
Conclusion: The current industrial production of profenols mainly relies on hydrolases. The bioreduction route established here represents an alternative method for the production of profenols that is competitive with hydrolase-catalyzed kinetic resolutions.
Keywords: Aldo–keto reductase engineering; Biocatalyst stability; Enantiopure 2-aryl-1-propanol; Reductive dynamic kinetic resolution.
© 2021. The Author(s).
Conflict of interest statement
The authors declare that they have no competing interests.
Figures
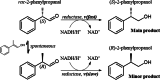
References
-
- Díaz-Rodríguez A, Ríos-Lombardía N, Sattler JH, Lavandera I, Gotor-Fernández V, Kroutil W, Gotor V. Deracemisation of profenol core by combining laccase/TEMPO-mediated oxidation and alcohol dehydrogenase-catalysed dynamic kinetic resolution. Catal Sci Technol. 2015;5:1443–1446. doi: 10.1039/C4CY01351D. - DOI
-
- Kourist R, Domínguez de María P, Miyamoto K. Biocatalytic strategies for the asymmetric synthesis of profens—recent trends and developments. Green Chem. 2011;13:2607–2618; 10.1039/C1GC15162B.
-
- Carvalho PO, Cass QB, Calafatti SA, Contesini FJ, Bizaco R. Review- Alternatives for the separation of drug enantiomers: ibuprofen as a model compound. Braz J Chem Eng. 2006;23:291–300. doi: 10.1590/S0104-66322006000300003. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

