Associations between alcohol use and accelerated biological ageing
- PMID: 34636470
- PMCID: PMC7614236
- DOI: 10.1111/adb.13100
Associations between alcohol use and accelerated biological ageing
Abstract
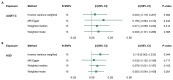
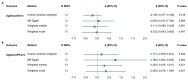
Harmful alcohol use is a leading cause of premature death and is associated with age-related disease. Biological ageing is highly variable between individuals and may deviate from chronological ageing, suggesting that biomarkers of biological ageing (derived from DNA methylation or brain structural measures) may be clinically relevant. Here, we investigated the relationships between alcohol phenotypes and both brain and DNA methylation age estimates. First, using data from UK Biobank and Generation Scotland, we tested the association between alcohol consumption (units/week) or hazardous use (Alcohol Use Disorders Identification Test [AUDIT] scores) and accelerated brain and epigenetic ageing in 20,258 and 8051 individuals, respectively. Second, we used Mendelian randomisation (MR) to test for a causal effect of alcohol consumption levels and alcohol use disorder (AUD) on biological ageing. Alcohol use showed a consistent positive association with higher predicted brain age (AUDIT-C: β = 0.053, p = 3.16 × 10-13 ; AUDIT-P: β = 0.052, p = 1.6 × 10-13 ; total AUDIT score: β = 0.062, p = 5.52 × 10-16 ; units/week: β = 0.078, p = 2.20 × 10-16 ), and two DNA methylation-based estimates of ageing, GrimAge (units/week: β = 0.053, p = 1.48 × 10-7 ) and PhenoAge (units/week: β = 0.077, p = 2.18x10-10 ). MR analyses revealed limited evidence for a causal effect of AUD on accelerated brain ageing (β = 0.118, p = 0.044). However, this result should be interpreted cautiously as the significant effect was driven by a single genetic variant. We found no evidence for a causal effect of alcohol consumption levels on accelerated biological ageing. Future studies investigating the mechanisms associating alcohol use with accelerated biological ageing are warranted.
Keywords: Generation Scotland; Mendelian randomisation; UK Biobank; alcohol use; brain ageing; epigenetic ageing.
© 2021 The Authors. Addiction Biology published by John Wiley & Sons Ltd on behalf of Society for the Study of Addiction.
Conflict of interest statement
AMM reports grants from The Sackler Trust, grants from Eli Lilly and grants from Janssen outside the submitted work. JHC is a shareholder in and scientific advisor to Brain Key, a medical image analysis software company. The remaining authors declare no conflicts of interest. This manuscript has been uploaded as a preprint to MedRxiv.
Figures



References
Publication types
MeSH terms
Grants and funding
- 104036/Z/14/Z/WT_/Wellcome Trust/United Kingdom
- 108890/Z/15/Z/WT_/Wellcome Trust/United Kingdom
- MC_QA137853/MRC_/Medical Research Council/United Kingdom
- 108890/WT_/Wellcome Trust/United Kingdom
- MC_UU_00011/3/MRC_/Medical Research Council/United Kingdom
- MR/R024065/1/MRC_/Medical Research Council/United Kingdom
- MC_PC_17228/MRC_/Medical Research Council/United Kingdom
- MR/R024790/2/MRC_/Medical Research Council/United Kingdom
- CZD/16/6/CSO_/Chief Scientist Office/United Kingdom
- BHF_/British Heart Foundation/United Kingdom
- 104036/WT_/Wellcome Trust/United Kingdom
- DH_/Department of Health/United Kingdom
- R01 AG054628/AG/NIA NIH HHS/United States
- CRUK_/Cancer Research UK/United Kingdom
- MC_UU_00011/1/MRC_/Medical Research Council/United Kingdom
LinkOut - more resources
Full Text Sources
Medical

