Adaptation of Bacillus thuringiensis to Plant Colonization Affects Differentiation and Toxicity
- PMID: 34636664
- PMCID: PMC8510532
- DOI: 10.1128/mSystems.00864-21
Adaptation of Bacillus thuringiensis to Plant Colonization Affects Differentiation and Toxicity
Abstract
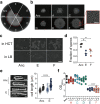
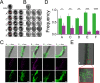
The Bacillus cereus group (Bacillus cereus sensu lato) has a diverse ecology, including various species that are vertebrate or invertebrate pathogens. Few isolates from the B. cereus group have however been demonstrated to benefit plant growth. Therefore, it is crucial to explore how bacterial development and pathogenesis evolve during plant colonization. Herein, we investigated Bacillus thuringiensis (Cry-) adaptation to the colonization of Arabidopsis thaliana roots and monitored changes in cellular differentiation in experimentally evolved isolates. Isolates from two populations displayed improved iterative ecesis on roots and increased virulence against insect larvae. Molecular dissection and recreation of a causative mutation revealed the importance of a nonsense mutation in the rho transcription terminator gene. Transcriptome analysis revealed how Rho impacts various B. thuringiensis genes involved in carbohydrate metabolism and virulence. Our work suggests that evolved multicellular aggregates have a fitness advantage over single cells when colonizing plants, creating a trade-off between swimming and multicellularity in evolved lineages, in addition to unrelated alterations in pathogenicity. IMPORTANCE Biologicals-based plant protection relies on the use of safe microbial strains. During application of biologicals to the rhizosphere, microbes adapt to the niche, including genetic mutations shaping the physiology of the cells. Here, the experimental evolution of Bacillus thuringiensis lacking the insecticide crystal toxins was examined on the plant root to reveal how adaptation shapes the differentiation of this bacterium. Interestingly, evolution of certain lineages led to increased hemolysis and insect larva pathogenesis in B. thuringiensis driven by transcriptional rewiring. Further, our detailed study reveals how inactivation of the transcription termination protein Rho promotes aggregation on the plant root in addition to altered differentiation and pathogenesis in B. thuringiensis.
Keywords: Arabidopsis thaliana; Bacillus thuringiensis; experimental evolution; pathogenesis; plant-microbe interaction.
Figures

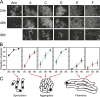



References
Grants and funding
LinkOut - more resources
Full Text Sources

